ROOT's RDataFrame offers a modern, high-level interface for analysis of data stored in TTree , CSV and other data formats, in C++ or Python.
You can use RDataFrame in Python thanks to the dynamic Python/C++ translation of PyROOT. In general, the interface is the same as for C++, a simple example follows. In the simple example that was shown above, a C++ expression is passed to the Filter() operation as a string ("x > 0"), even if we call the method from Python. Indeed, under the hood, the analysis computations run in C++, while Python is just the interface language. To perform more complex operations that don't fit into a simple expression string, you can just-in-time compile C++ functions - via the C++ interpreter cling - and use those functions in an expression. See the following snippet for an example: To increase the performance even further, you can also pre-compile a C++ library with full code optimizations and load the function into the RDataFrame computation as follows. A more thorough explanation of how to use C++ code from Python can be found in the PyROOT manual. ROOT also offers the option to compile Python functions with fundamental types and arrays thereof using Numba. Such compiled functions can then be used in a C++ expression provided to RDataFrame. The function to be compiled should be decorated with ROOT.Numba.Declare, which allows to specify the parameter and return types. See the following snippet for a simple example or the full tutorial here. It also works with collections: RVec objects of fundamental types can be transparently converted to/from numpy arrays: Note that this functionality requires the Python packages numba and cffi to be installed. Eventually, you probably would like to inspect the content of the RDataFrame or process the data further with Python libraries. For this purpose, we provide the AsNumpy() function, which returns the columns of your RDataFrame as a dictionary of NumPy arrays. See a few simple examples below or a full tutorial here. If your column contains scalar values of fundamental types (e.g., integers, floats), AsNumpy() produces NumPy arrays with the appropriate dtype: Columns containing non-fundamental types (e.g., objects, strings) will result in NumPy arrays with dtype=object. If your column contains collections of fundamental types (e.g., std::vector<int>), AsNumpy() produces a NumPy array with dtype=object where each element is a NumPy array representing the collection for its corresponding entry in the column. If the collection at a certain entry contains values of fundamental types, or if it is a regularly shaped multi-dimensional array of a fundamental type, then the numpy array representing the collection for that entry will have the dtype associated with the value type of the collection, for example: If the collection at a certain entry contains values of a non-fundamental type, AsNumpy() will fallback on the default behavior and produce a NumPy array with dtype=object for that collection. For more complex collection types in your entries, e.g. when every entry has a jagged array value, refer to the section on interoperability with AwkwardArray. In case you have data in NumPy arrays in Python and you want to process the data with ROOT, you can easily create an RDataFrame using ROOT.RDF.FromNumpy. The factory function accepts a dictionary where the keys are the column names and the values are NumPy arrays, and returns a new RDataFrame with the provided columns. Only arrays of fundamental types (integers and floating point values) are supported and the arrays must have the same length. Data is read directly from the arrays: no copies are performed. The function for RDataFrame to Awkward conversion is ak.from_rdataframe(). The argument to this function accepts a tuple of strings that are the RDataFrame column names. By default this function returns ak.Array type. The function for Awkward to RDataFrame conversion is ak.to_rdataframe(). The argument to this function requires a dictionary: { <column name string> : <awkward array> }. This function always returns an RDataFrame object. The arrays given for each column have to be equal length: The Histo1D(), Histo2D(), Histo3D(), Profile1D() and Profile2D() methods return histograms and profiles, respectively, which can be constructed using a model argument. In Python, we can specify the arguments for the constructor of such histogram or profile model with a Python tuple, as shown in the example below: The ROOT::RDF::AsRNode function casts an RDataFrame node to the generic ROOT::RDF::RNode type. From Python, it can be used to pass any RDataFrame node as an argument of a C++ function, as shown below:
Python interface
User code in the RDataFrame workflow
C++ code
Python code
Interoperability with NumPy
Conversion to NumPy arrays
Scalar columns
Collection Columns
Processing data stored in NumPy arrays
Interoperability with AwkwardArray
Construct histogram and profile models from a tuple
AsRNode helper function
In addition, multi-threading and other low-level optimisations allow users to exploit all the resources available on their machines completely transparently.
Skip to the class reference or keep reading for the user guide.
In a nutshell:
Calculations are expressed in terms of a type-safe functional chain of actions and transformations, RDataFrame takes care of their execution. The implementation automatically puts in place several low level optimisations such as multi-thread parallelization and caching.
You can directly see RDataFrame in action in our tutorials, in C++ or Python.
These are the operations which can be performed with RDataFrame.
Transformations are a way to manipulate the data.
| Transformation | Description |
|---|---|
| Alias() | Introduce an alias for a particular column name. |
| DefaultValueFor() | If the value of the input column is missing, provide a default value instead. |
| Define() | Create a new column in the dataset. Example usages include adding a column that contains the invariant mass of a particle, or a selection of elements of an array (e.g. only the pts of "good" muons). |
| DefinePerSample() | Define a new column that is updated when the input sample changes, e.g. when switching tree being processed in a chain. |
| DefineSlot() | Same as Define(), but the user-defined function must take an extra unsigned int slot as its first parameter. slot will take a different value, in the range [0, nThread-1], for each thread of execution. This is meant as a helper in writing thread-safe Define() transformations when using RDataFrame after ROOT::EnableImplicitMT(). DefineSlot() works just as well with single-thread execution: in that case slot will always be 0. |
| DefineSlotEntry() | Same as DefineSlot(), but the entry number is passed in addition to the slot number. This is meant as a helper in case the expression depends on the entry number. For details about entry numbers in multi-threaded runs, see here. |
| Filter() | Filter rows based on user-defined conditions. |
| FilterAvailable() | Specialized Filter. If the value of the input column is available, keep the entry, otherwise discard it. |
| FilterMissing() | Specialized Filter. If the value of the input column is missing, keep the entry, otherwise discard it. |
| Range() | Filter rows based on entry number (single-thread only). |
| Redefine() | Overwrite the value and/or type of an existing column. See Define() for more information. |
| RedefineSlot() | Overwrite the value and/or type of an existing column. See DefineSlot() for more information. |
| RedefineSlotEntry() | Overwrite the value and/or type of an existing column. See DefineSlotEntry() for more information. |
| Vary() | Register systematic variations for an existing column. Varied results are then extracted via VariationsFor(). |
Actions aggregate data into a result. Each one is described in more detail in the reference guide.
In the following, whenever we say an action "returns" something, we always mean it returns a smart pointer to it. Actions only act on events that pass all preceding filters.
Lazy actions only trigger the event loop when one of the results is accessed for the first time, making it easy to produce many different results in one event loop. Instant actions trigger the event loop instantly.
| Lazy action | Description |
|---|---|
| Aggregate() | Execute a user-defined accumulation operation on the processed column values. |
| Book() | Book execution of a custom action using a user-defined helper object. |
| Cache() | Cache column values in memory. Custom columns can be cached as well, filtered entries are not cached. Users can specify which columns to save (default is all). |
| Count() | Return the number of events processed. Useful e.g. to get a quick count of the number of events passing a Filter. |
| Display() | Provides a printable representation of the dataset contents. The method returns a ROOT::RDF::RDisplay() instance which can print a tabular representation of the data or return it as a string. |
| Fill() | Fill a user-defined object with the values of the specified columns, as if by calling Obj.Fill(col1, col2, ...). |
| Graph() | Fills a TGraph with the two columns provided. If multi-threading is enabled, the order of the points may not be the one expected, it is therefore suggested to sort if before drawing. |
| GraphAsymmErrors() | Fills a TGraphAsymmErrors. Should be used for any type of graph with errors, including cases with errors on one of the axes only. If multi-threading is enabled, the order of the points may not be the one expected, it is therefore suggested to sort if before drawing. |
| Histo1D(), Histo2D(), Histo3D() | Fill a one-, two-, three-dimensional histogram with the processed column values. |
| HistoND() | Fill an N-dimensional histogram with the processed column values. |
| HistoNSparseD() | Fill an N-dimensional sparse histogram with the processed column values. Memory is allocated only for non-empty bins. |
| Max() | Return the maximum of processed column values. If the type of the column is inferred, the return type is double, the type of the column otherwise. |
| Mean() | Return the mean of processed column values. |
| Min() | Return the minimum of processed column values. If the type of the column is inferred, the return type is double, the type of the column otherwise. |
| Profile1D(), Profile2D() | Fill a one- or two-dimensional profile with the column values that passed all filters. |
| Reduce() | Reduce (e.g. sum, merge) entries using the function (lambda, functor...) passed as argument. The function must have signature T(T,T) where T is the type of the column. Return the final result of the reduction operation. An optional parameter allows initialization of the result object to non-default values. |
| Report() | Obtain statistics on how many entries have been accepted and rejected by the filters. See the section on named filters for a more detailed explanation. The method returns a ROOT::RDF::RCutFlowReport instance which can be queried programmatically to get information about the effects of the individual cuts. |
| Stats() | Return a TStatistic object filled with the input columns. |
| StdDev() | Return the unbiased standard deviation of the processed column values. |
| Sum() | Return the sum of the values in the column. If the type of the column is inferred, the return type is double, the type of the column otherwise. |
| Take() | Extract a column from the dataset as a collection of values, e.g. a std::vector<float> for a column of type float. |
| Instant action | Description |
|---|---|
| Foreach() | Execute a user-defined function on each entry. Users are responsible for the thread-safety of this callable when executing with implicit multi-threading enabled. |
| ForeachSlot() | Same as Foreach(), but the user-defined function must take an extra unsigned int slot as its first parameter. slot will take a different value, 0 to nThreads - 1, for each thread of execution. This is meant as a helper in writing thread-safe Foreach() actions when using RDataFrame after ROOT::EnableImplicitMT(). ForeachSlot() works just as well with single-thread execution: in that case slot will always be 0. |
| Snapshot() | Write the processed dataset to disk, in a new TTree or RNTuple and TFile. Custom columns can be saved as well, filtered entries are not saved. Users can specify which columns to save (default is all nominal columns). Columns resulting from Vary() can be included by setting the corresponding flag in RSnapshotOptions. Snapshot, by default, overwrites the output file if it already exists. Snapshot() can be made lazy setting the appropriate flag in the snapshot options. |
These operations do not modify the dataframe or book computations but simply return information on the RDataFrame object.
| Operation | Description |
|---|---|
| Describe() | Get useful information describing the dataframe, e.g. columns and their types. |
| GetColumnNames() | Get the names of all the available columns of the dataset. |
| GetColumnType() | Return the type of a given column as a string. |
| GetColumnTypeNamesList() | Return the list of type names of columns in the dataset. |
| GetDefinedColumnNames() | Get the names of all the defined columns. |
| GetFilterNames() | Return the names of all filters in the computation graph. |
| GetNRuns() | Return the number of event loops run by this RDataFrame instance so far. |
| GetNSlots() | Return the number of processing slots that RDataFrame will use during the event loop (i.e. the concurrency level). |
| ROOT::RDF::SaveGraph() | Store the computation graph of an RDataFrame in DOT format (graphviz) for easy inspection. See the relevant section for details. |
Users define their analysis as a sequence of operations to be performed on the dataframe object; the framework takes care of the management of the loop over entries as well as low-level details such as I/O and parallelization. RDataFrame provides methods to perform most common operations required by ROOT analyses; at the same time, users can just as easily specify custom code that will be executed in the event loop.
RDataFrame is built with a modular and flexible workflow in mind, summarised as follows:
Make sure to book all transformations and actions before you access the contents of any of the results. This lets RDataFrame accumulate work and then produce all results at the same time, upon first access to any of them.
The following table shows how analyses based on TTreeReader and TTree::Draw() translate to RDataFrame. Follow the crash course to discover more idiomatic and flexible ways to express analyses with RDataFrame.
| TTreeReader | ROOT::RDataFrame |
TTreeReader reader("myTree", file);
while(reader.Next()) {
}
An interface for reading values stored in ROOT columnar datasets. Definition TTreeReaderValue.h:205 A simple, robust and fast interface to read values from ROOT columnar datasets such as TTree,... Definition TTreeReader.h:46 | d.Filter(IsGoodEvent).Foreach(DoStuff);
|
| TTree::Draw | ROOT::RDataFrame |
ROOT::RDataFrame df("myTree", file);
h->Draw()
| |
tree->Draw("jet_eta", "weight*(event == 1)");
| df.Filter("event == 1").Histo1D("jet_eta", "weight");
// or the fully compiled version:
df.Filter([] (ULong64_t e) { return e == 1; }, {"event"}).Histo1D<RVec<float>>("jet_eta", "weight");
RResultPtr<::TH1D > Histo1D(const TH1DModel &model={"", "", 128u, 0., 0.}, std::string_view vName="") Definition RInterface.hxx:1978 |
// object selection: for each event, fill histogram with array of selected pts
tree->Draw('Muon_pt', 'Muon_pt > 100');
| // with RDF, arrays are read as ROOT::VecOps::RVec objects
df.Define("good_pt", "Muon_pt[Muon_pt > 100]").Histo1D("good_pt")
|
All snippets of code presented in the crash course can be executed in the ROOT interpreter. Simply precede them with
which is omitted for brevity. The terms "column" and "branch" are used interchangeably.
RDataFrame's constructor is where the user specifies the dataset and, optionally, a default set of columns that operations should work with. Here are the most common methods to construct an RDataFrame object:
Additionally, users can construct an RDataFrame with no data source by passing an integer number. This is the number of rows that will be generated by this RDataFrame.
This is useful to generate simple datasets on the fly: the contents of each event can be specified with Define() (explained below). For example, we have used this method to generate Pythia events and write them to disk in parallel (with the Snapshot action).
For data sources other than TTrees and TChains, RDataFrame objects are constructed using ad-hoc factory functions (see e.g. FromCSV(), FromSqlite(), FromArrow()):
Let's now tackle a very common task, filling a histogram:
The first line creates an RDataFrame associated to the TTree "myTree". This tree has a branch named "MET".
Histo1D() is an action; it returns a smart pointer (a ROOT::RDF::RResultPtr, to be precise) to a TH1D histogram filled with the MET of all events. If the quantity stored in the column is a collection (e.g. a vector or an array), the histogram is filled with all vector elements for each event.
You can use the objects returned by actions as if they were pointers to the desired results. There are many other possible actions, and all their results are wrapped in smart pointers; we'll see why in a minute.
Let's say we want to cut over the value of branch "MET" and count how many events pass this cut. This is one way to do it:
The filter string (which must contain a valid C++ expression) is applied to the specified columns for each event; the name and types of the columns are inferred automatically. The string expression is required to return a bool which signals whether the event passes the filter (true) or not (false).
You can think of your data as "flowing" through the chain of calls, being transformed, filtered and finally used to perform actions. Multiple Filter() calls can be chained one after another.
Using string filters is nice for simple things, but they are limited to specifying the equivalent of a single return statement or the body of a lambda, so it's cumbersome to use strings with more complex filters. They also add a small runtime overhead, as ROOT needs to just-in-time compile the string into C++ code. When more freedom is required or runtime performance is very important, a C++ callable can be specified instead (a lambda in the following snippet, but it can be any kind of function or even a functor class), together with a list of column names. This snippet is analogous to the one above:
An example of a more complex filter expressed as a string containing C++ code is shown below
The code snippet above defines a column p that is a fixed-size array using the component column names and then filters on its magnitude by looping over its elements. It must be noted that the usage of strings to define columns like the one above is currently the only possibility when using PyROOT. When writing expressions as such, only constants and data coming from other columns in the dataset can be involved in the code passed as a string. Local variables and functions cannot be used, since the interpreter will not know how to find them. When capturing local state is necessary, it must first be declared to the ROOT C++ interpreter.
More information on filters and how to use them to automatically generate cutflow reports can be found below.
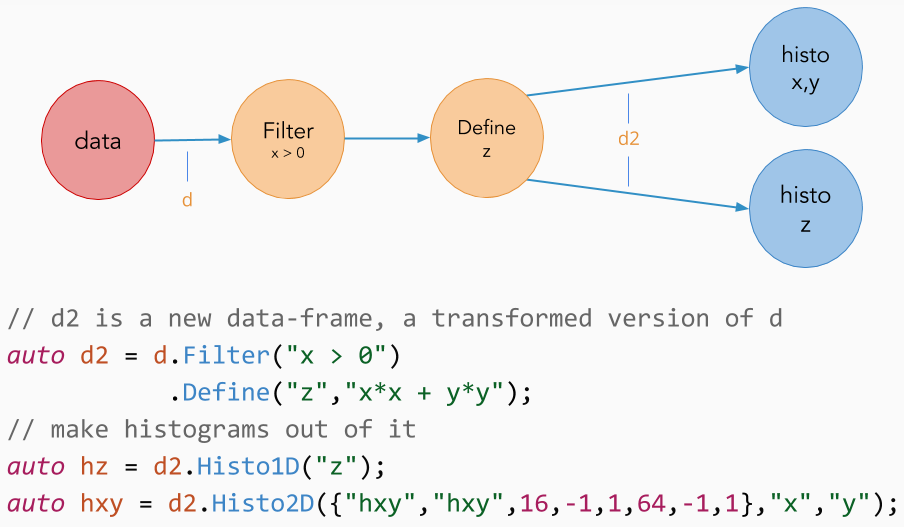
Let's now consider the case in which "myTree" contains two quantities "x" and "y", but our analysis relies on a derived quantity z = sqrt(x*x + y*y). Using the Define() transformation, we can create a new column in the dataset containing the variable "z":
Define() creates the variable "z" by applying sqrtSum to "x" and "y". Later in the chain of calls we refer to variables created with Define() as if they were actual tree branches/columns, but they are evaluated on demand, at most once per event. As with filters, Define() calls can be chained with other transformations to create multiple custom columns. Define() and Filter() transformations can be concatenated and intermixed at will.
As with filters, it is possible to specify new columns as string expressions. This snippet is analogous to the one above:
Again the names of the columns used in the expression and their types are inferred automatically. The string must be valid C++ and it is just-in-time compiled. The process has a small runtime overhead and like with filters it is currently the only possible approach when using PyROOT.
Previously, when showing the different ways an RDataFrame can be created, we showed a constructor that takes a number of entries as a parameter. In the following example we show how to combine such an "empty" RDataFrame with Define() transformations to create a dataset on the fly. We then save the generated data on disk using the Snapshot() action.
This example is slightly more advanced than what we have seen so far. First, it makes use of lambda captures (a simple way to make external variables available inside the body of C++ lambdas) to act on the same variable x from both Define() transformations. Second, we have stored the transformed dataframe in a variable. This is always possible, since at each point of the transformation chain users can store the status of the dataframe for further use (more on this below).
You can read more about defining new columns here.
It is sometimes necessary to limit the processing of the dataset to a range of entries. For this reason, the RDataFrame offers the concept of ranges as a node of the RDataFrame chain of transformations; this means that filters, columns and actions can be concatenated to and intermixed with Range()s. If a range is specified after a filter, the range will act exclusively on the entries passing the filter – it will not even count the other entries! The same goes for a Range() hanging from another Range(). Here are some commented examples:
Note that ranges are not available when multi-threading is enabled. More information on ranges is available here.
As a final example let us apply two different cuts on branch "MET" and fill two different histograms with the "pt_v" of the filtered events. By now, you should be able to easily understand what is happening:
RDataFrame executes all above actions by running the event-loop only once. The trick is that actions are not executed at the moment they are called, but they are lazy, i.e. delayed until the moment one of their results is accessed through the smart pointer. At that time, the event loop is triggered and all results are produced simultaneously.
For yet another example of the difference between the correct and incorrect running of the event-loop, see the following two code snippets. We assume our ROOT file has branches a, b and c.
The correct way - the dataset is only processed once.
An incorrect way - the dataset is processed three times.
It is therefore good practice to declare all your transformations and actions before accessing their results, allowing RDataFrame to run the loop once and produce all results in one go.
Let's say we would like to run the previous examples in parallel on several cores, dividing events fairly between cores. The only modification required to the snippets would be the addition of this line before constructing the main dataframe object:
Simple as that. More details are given below.
RDataFrame reads collections as the special type ROOT::RVec: for example, a column containing an array of floating point numbers can be read as a ROOT::RVecF. C-style arrays (with variable or static size), STL vectors and most other collection types can be read this way.
RVec is a container similar to std::vector (and can be used just like a std::vector) but it also offers a rich interface to operate on the array elements in a vectorised fashion, similarly to Python's NumPy arrays.
For example, to fill a histogram with the "pt" of selected particles for each event, Define() can be used to create a column that contains the desired array elements as follows:
And in Python:
Learn more at ROOT::VecOps::RVec.
ROOT provides convenience utility functions to work with RVec which are available in the ROOT::VecOps namespace
A filter is created through a call to Filter(f, columnList) or Filter(filterString). In the first overload, f can be a function, a lambda expression, a functor class, or any other callable object. It must return a bool signalling whether the event has passed the selection (true) or not (false). It should perform "read-only" operations on the columns, and should not have side-effects (e.g. modification of an external or static variable) to ensure correctness when implicit multi-threading is active. The second overload takes a string with a valid C++ expression in which column names are used as variable names (e.g. Filter("x[0] + x[1] > 0")). This is a convenience feature that comes with a certain runtime overhead: C++ code has to be generated on the fly from this expression before using it in the event loop. See the paragraph about "Just-in-time compilation" below for more information.
RDataFrame only evaluates filters when necessary: if multiple filters are chained one after another, they are executed in order and the first one returning false causes the event to be discarded and triggers the processing of the next entry. If multiple actions or transformations depend on the same filter, that filter is not executed multiple times for each entry: after the first access it simply serves a cached result.
An optional string parameter name can be passed to the Filter() method to create a named filter. Named filters work as usual, but also keep track of how many entries they accept and reject.
Statistics are retrieved through a call to the Report() method:
Stats are stored in the same order as named filters have been added to the graph, and refer to the latest event-loop that has been run using the relevant RDataFrame.
When RDataFrame is not being used in a multi-thread environment (i.e. no call to EnableImplicitMT() was made), Range() transformations are available. These act very much like filters but instead of basing their decision on a filter expression, they rely on begin,end and stride parameters.
The actual number of entries processed downstream of a Range() node will be (end - begin)/stride (or less if less entries than that are available).
Note that ranges act "locally", not based on the global entry count: Range(10,50) means "skip the first 10 entries that reach this node*, let the next 40 entries pass, then stop processing". If a range node hangs from a filter node, and the range has a begin parameter of 10, that means the range will skip the first 10 entries that pass the preceding filter.
Ranges allow "early quitting": if all branches of execution of a functional graph reached their end value of processed entries, the event-loop is immediately interrupted. This is useful for debugging and quick data explorations.
Custom columns are created by invoking Define(name, f, columnList). As usual, f can be any callable object (function, lambda expression, functor class...); it takes the values of the columns listed in columnList (a list of strings) as parameters, in the same order as they are listed in columnList. f must return the value that will be assigned to the temporary column.
A new variable is created called name, accessible as if it was contained in the dataset from subsequent transformations/actions.
Use cases include:
An exception is thrown if the name of the new column/branch is already in use for another branch in the TTree.
It is also possible to specify the quantity to be stored in the new temporary column as a C++ expression with the method Define(name, expression). For example this invocation
will create a new column called "pt" the value of which is calculated starting from the columns px and py. The system builds a just-in-time compiled function starting from the expression after having deduced the list of necessary branches from the names of the variables specified by the user.
It is possible to create custom columns also as a function of the processing slot and entry numbers. The methods that can be invoked are:
Actions can be instant or lazy. Instant actions are executed as soon as they are called, while lazy actions are executed whenever the object they return is accessed for the first time. As a rule of thumb, actions with a return value are lazy, the others are instant.
When a lazy action is called, it returns a ROOT::RDF::RResultPtr<T>, where T is the type of the result of the action. The final result will be stored in the RResultPtr, and can be retrieved by dereferencing it or via its GetValue method. Retrieving the result also starts the event loop if the result hasn't been produced yet.
The RResultPtr shares ownership of the result object. To directly access result, use:
To return results from functions, a copy of the underlying shared_ptr can be obtained:
If the result had been returned by reference or bare pointer, it would have gotten destroyed when the function exits.
To share ownership but not produce the result ("keep it lazy"), copy the RResultPtr:
If the type of the return value of an action is a collection, e.g. std::vector<int>, you can iterate its elements directly through the wrapping RResultPtr:
An action that needs values for its computations will request it from a reader, e.g. a column created via Define or available from the input dataset. The action will request values from each column of the list of input columns (either inferred or specified by the user), in order. For example:
The Graph action is going to request first the value from column "x", then that of column "y". Specifically, the order of execution of the operations of nodes in this branch of the computation graph is guaranteed to be top to bottom.
RDataFrame applications can be executed in parallel through distributed computing frameworks on a set of remote machines thanks to the Python package ROOT.RDF.Distributed. This Python-only package allows to scale the optimized performance RDataFrame can achieve on a single machine to multiple nodes at the same time. It is designed so that different backends can be easily plugged in, currently supporting Apache Spark and Dask. Here is a minimal example usage of distributed RDataFrame:
The main goal of this package is to support running any RDataFrame application distributedly. Nonetheless, not all parts of the RDataFrame API currently work with this package. The subset that is currently available is:
with support for more operations coming in the future. Currently, to the supported data sources belong TTree, TChain, RNTuple and RDatasetSpec.
In order to distribute the RDataFrame workload, you can connect to a Spark cluster you have access to through the official Spark API, then hook the connection instance to the distributed RDataFrame object like so:
Note that with the usage above the case of executor = None is not supported. One can explicitly create a ROOT.RDF.Distributed.Spark.RDataFrame object in order to get a default instance of SparkContext in case it is not already provided as argument.
Similarly, you can connect to a Dask cluster by creating your own connection object which internally operates with one of the cluster schedulers supported by Dask (more information in the Dask distributed docs):
Note that with the usage above the case of executor = None is not supported. One can explicitly create a ROOT.RDF.Distributed.Dask.RDataFrame object in order to get a default instance of distributed.Client in case it is not already provided as argument. This will run multiple processes on the local machine using all available cores.
A distributed RDataFrame has internal logic to define in how many chunks the input dataset will be split before sending tasks to the distributed backend. Each task reads and processes one of said chunks. The logic is backend-dependent, but generically tries to infer how many cores are available in the cluster through the connection object. The number of tasks will be equal to the inferred number of cores. There are cases where the connection object of the chosen backend doesn't have information about the actual resources of the cluster. An example of this is when using Dask to connect to a batch system. The client object created at the beginning of the application does not automatically know how many cores will be available during distributed execution, since the jobs are submitted to the batch system after the creation of the connection. In such cases, the logic is to default to process the whole dataset in 2 tasks.
The number of tasks submitted for distributed execution can be also set programmatically, by providing the optional keyword argument npartitions when creating the RDataFrame object. This parameter is accepted irrespectively of the backend used:
Note that when processing a TTree or TChain dataset, the npartitions value should not exceed the number of clusters in the dataset. The number of clusters in a TTree can be retrieved by typing rootls -lt myfile.root at a command line.
RDataFrame can be also built from a JSON sample specification file using the FromSpec function. In distributed mode, two arguments need to be provided: the path to the specification jsonFile (same as for local RDF case) and an additional executor argument - in the same manner as for the RDataFrame constructors above - an executor can either be a spark connection or a dask client. If no second argument is given, the local version of FromSpec will be run. Here is an example of FromSpec usage in distributed RDF using either spark or dask backends. For more information on FromSpec functionality itself please refer to FromSpec documentation. Note that adding metadata and friend information is supported, but adding the global range will not be respected in the distributed execution.
Using spark:
Using dask:
The Snapshot operation behaves slightly differently when executed distributedly. First off, it requires the path supplied to the Snapshot call to be accessible from any worker of the cluster and from the client machine (in general it should be provided as an absolute path). Another important difference is that n separate files will be produced, where n is the number of dataset partitions. As with local RDataFrame, the result of a Snapshot on a distributed RDataFrame is another distributed RDataFrame on which we can define a new computation graph and run more distributed computations.
Submitting multiple distributed RDataFrame executions is supported through the RunGraphs function. Similarly to its local counterpart, the function expects an iterable of objects representing an RDataFrame action. Each action will be triggered concurrently to send multiple computation graphs to a distributed cluster at the same time:
Every distributed backend supports this feature and graphs belonging to different backends can be still triggered with a single call to RunGraphs (e.g. it is possible to send a Spark job and a Dask job at the same time).
When calling a Histo*D operation in distributed mode, remember to pass to the function the model of the histogram to be computed, e.g. the axis range and the number of bins:
Without this, two partial histograms resulting from two distributed tasks would have incompatible binning, thus leading to errors when merging them. Failing to pass a histogram model will raise an error on the client side, before starting the distributed execution.
The live visualization feature allows real-time data representation of plots generated during the execution of a distributed RDataFrame application. It enables visualizing intermediate results as they are computed across multiple nodes of a Dask cluster by creating a canvas and continuously updating it as partial results become available.
The LiveVisualize() function can be imported from the Python package ROOT.RDF.Distributed:
The function takes drawable objects (e.g. histograms) and optional callback functions as argument, it accepts 4 different input formats:
Distributed RDF provides an interface for the users who want to inject the C++ code (via header files, shared libraries or declare the code directly) into their distributed RDF application, or their application needs to use information from external files which should be distributed to the workers (for example, a JSON or a txt file with necessary parameters information).
The examples below show the usage of these interface functions: firstly, how this is done in a local Python RDF application and secondly, how it is done distributedly.
The cpp code is always available to all dataframes.
As pointed out before in this document, RDataFrame can transparently perform multi-threaded event loops to speed up the execution of its actions. Users have to call ROOT::EnableImplicitMT() before constructing the RDataFrame object to indicate that it should take advantage of a pool of worker threads. Each worker thread processes a distinct subset of entries, and their partial results are merged before returning the final values to the user.
By default, RDataFrame will use as many threads as the hardware supports, using up all the resources on a machine. This might be undesirable on shared computing resources such as a batch cluster. Therefore, when running on shared computing resources, use
or export an environment variable:
replacing numThreads with the number of CPUs/slots that were allocated for this job.
RDataFrame operations such as Histo1D() or Snapshot() are guaranteed to work correctly in multi-thread event loops. User-defined expressions, such as strings or lambdas passed to Filter(), Define(), Foreach(), Reduce() or Aggregate() will have to be thread-safe, i.e. it should be possible to call them concurrently from different threads.
Note that simple Filter() and Define() transformations will inherently satisfy this requirement: Filter() / Define() expressions will often be pure in the functional programming sense (no side-effects, no dependency on external state), which eliminates all risks of race conditions.
In order to facilitate writing of thread-safe operations, some RDataFrame features such as Foreach(), Define() or OnPartialResult() offer thread-aware counterparts (ForeachSlot(), DefineSlot(), OnPartialResultSlot()): their only difference is that they will pass an extra slot argument (an unsigned integer) to the user-defined expression. When calling user-defined code concurrently, RDataFrame guarantees that different threads will see different values of the slot parameter, where slot will be a number between 0 and GetNSlots() - 1. Note that not all slot numbers may be reached, or some slots may be reached more often depending on how computation tasks are scheduled. In other words, within a slot, computations run sequentially, and events are processed sequentially. Note that the same slot might be associated to different threads over the course of a single event loop, but two threads will never receive the same slot at the same time. This extra parameter might facilitate writing safe parallel code by having each thread write/modify a different processing slot, e.g. a different element of a list. See here for an example usage of ForeachSlot().
A complex analysis may require multiple separate RDataFrame computation graphs to produce all desired results. This poses the challenge that the event loops of each computation graph can be parallelized, but the different loops run sequentially, one after the other. On many-core architectures it might be desirable to run different event loops concurrently to improve resource usage. ROOT::RDF::RunGraphs() allows running multiple RDataFrame event loops concurrently:
To obtain the maximum performance out of RDataFrame, make sure to avoid just-in-time compiled versions of transformations and actions if at all possible. For instance, Filter("x > 0") requires just-in-time compilation of the corresponding C++ logic, while the equivalent Filter([](float x) { return x > 0.; }, {"x"}) does not. Similarly, Histo1D("x") requires just-in-time compilation after the type of x is retrieved from the dataset, while Histo1D<float>("x") does not; the latter spelling should be preferred for performance-critical applications.
Python applications cannot easily specify template parameters or pass C++ callables to RDataFrame. See Python interface for possible ways to speed up hot paths in this case.
Just-in-time compilation happens once, right before starting an event loop. To reduce the runtime cost of this step, make sure to book all operations for all RDataFrame computation graphs before the first event loop is triggered: just-in-time compilation will happen once for all code required to be generated up to that point, also across different computation graphs.
Also make sure not to count the just-in-time compilation time (which happens once before the event loop and does not depend on the size of the dataset) as part of the event loop runtime (which scales with the size of the dataset). RDataFrame has an experimental logging feature that simplifies measuring the time spent in just-in-time compilation and in the event loop (as well as providing some more interesting information). See Activating RDataFrame execution logs.
There are two reasons why RDataFrame may consume more memory than expected.
In multithreaded runs, each worker thread will create a local copy of histograms, which e.g. in case of many (possibly multi-dimensional) histograms with fine binning can result in significant memory consumption during the event loop. The thread-local copies of the results are destroyed when the final result is produced. Reducing the number of threads or using coarser binning will reduce the memory usage. For three-dimensional histograms, the number of clones can be reduced using ROOT::RDF::Experimental::ThreadsPerTH3().
When TH3s are shared among threads, TH3D will either be filled under lock (slowing down the execution) or using atomics if C++20 is available. The latter is significantly faster. The best value for ThreadsPerTH3 depends on the computation graph that runs. Use lower numbers such as 4 for speed and higher memory consumption, and higher numbers such as 16 for slower execution and memory savings.
Secondly, just-in-time compilation of string expressions or non-templated actions (see the previous paragraph) causes Cling, ROOT's C++ interpreter, to allocate some memory for the generated code that is only released at the end of the application. This commonly results in memory usage creep in long-running applications that create many RDataFrames one after the other. Possible mitigations include creating and running each RDataFrame event loop in a sub-process, or booking all operations for all different RDataFrame computation graphs before the first event loop is triggered, so that the interpreter is invoked only once for all computation graphs:
Here is a list of the most important features that have been omitted in the "Crash course" for brevity. You don't need to read all these to start using RDataFrame, but they are useful to save typing time and runtime.
Starting from ROOT v6.26, RDataFrame provides a flexible syntax to define systematic variations. This is done in two steps: a) register variations for one or more existing columns using Vary() and b) extract variations of normal RDataFrame results using VariationsFor(). In between these steps, no other change to the analysis code is required: the presence of systematic variations for certain columns is automatically propagated through filters, defines and actions, and RDataFrame will take these dependencies into account when producing varied results. VariationsFor() is included in header ROOT/RDFHelpers.hxx. The compiled C++ programs must include this header explicitly, this is not required for ROOT macros.
An example usage of Vary() and VariationsFor() in C++:
A shorter expression syntax is allowed for convenience (see the docs of the Vary overloads for more details):
A list of variation "tags" is passed as the last argument to Vary(). The tags give names to the varied values that are returned as elements of an RVec of the appropriate C++ type. The number of variation tags must correspond to the number of elements of this RVec (2 in the example above: the first element will correspond to the tag "down", the second to the tag "up"). The full variation name will be composed of the varied column name and the variation tags (e.g. "pt:down", "pt:up" in this example). Python usage looks similar.
Note how we use the "pt" column as usual in the Filter() and Define() calls and we simply use "x" as the value to fill the resulting histogram. To produce the varied results, RDataFrame will automatically execute the Filter and Define calls for each variation and fill the histogram with values and cuts that depend on the variation.
There is no limitation to the complexity of a Vary() expression. Just like for the Define() and Filter() calls, users are not limited to string expressions but they can also pass any valid C++ callable, including lambda functions and complex functors. The callable can be applied to zero or more existing columns and it will always receive their nominal value in input.
See the Vary() method for more information and this tutorial for an example usage of Vary and VariationsFor() in the analysis.
To combine Vary() and Snapshot(), the fIncludeVariations flag in RSnapshotOptions has to be set, as demonstrated in the following example:
This snapshot action will create an output file with a dataset that includes the columns requested in the Snapshot() call and their corresponding variations:
| Row | x | y | x__xVar_0 | y__xVar_0 | x__xVar_1 | y__xVar_1 | R_rdf_mask_t_0 |
|---|---|---|---|---|---|---|---|
| 0 | 0 | -0 | -0.5 | 0.5 | 0.5 | -0.5 | 111 |
| 1 | 10 | -10 | 9.5 | -9.5 | 10.5 | -10.5 | 111 |
| 2 | 20 | -20 | 19.5 | -19.5 | 20.5 | -20.5 | 111 |
| 3 | 30 | -30 | 29.5 | -29.5 | 30.5 | -30.5 | 111 |
| 4 | 40 | -40 | 39.5 | -39.5 | 40.5 | -40.5 | 111 |
| 5 | 0 | 0 | 49.5 | -49.5 | 0 | 0 | 010 |
| 6 | * Not written * | ||||||
| 7 | 0 | 0 | 0 | 0 | 70.5 | -70.5 | 100 |
"x" and "y" are the nominal values, and their variations are snapshot as <ColumnName>__<VariationName>_<VariationTag>.
When Filters are employed, some variations might not pass the selection cuts (like the rows 5 and 7 in the table above). In that case, RDataFrame will snapshot the filtered columns in a memory-efficient way by writing zero into the memory of fundamental types, or write a default-constructed object in case of classes. If none of the filters pass like in row 6, the entire event is omitted from the snapshot.
To tell apart a genuine 0 (like x in row 0) from a case where nominal or variation didn't pass a selection, RDataFrame writes a bitmask for each event, see last column of the table above. Every bit indicates whether its associated columns are valid. The bitmask is implemented as a 64-bit std::bitset in memory, written to the output dataset as a std::uin64_t. For every 64 columns, a new bitmask column is added to the output dataset.
For each column that gets varied, the nominal and all variation columns are each assigned a bit to denote whether their entries are valid. A mapping of column names to the corresponding bitmask is placed in the same file as the output dataset, with a name that follows the pattern "R_rdf_column_to_bitmask_mapping_<NAME_OF_THE_DATASET>". It is of type std::unordered_map<std::string, std::pair<std::string, unsigned int>>, and maps a column name to the name of the bitmask column and the index of the relevant bit. For example, in the same file as the dataset "Events" there would be an object named R_rdf_column_to_bitmask_mapping_Events. This object for example would describe a connection such as:
which means that the validity of the entries in muon_pt is established by the bit 42 in the bitmask found in the column R_rdf_mask_Events_0.
When RDataFrame opens a file, it checks for the existence of this mapping between columns and bitmasks, and loads it automatically if found. As such, RDataFrame makes the treatment of the various bitmap maskings completely transparent to the user.
In case certain values are labeled invalid by the corresponding bit, this will result in reading a missing value. The semantics of such a scenario follow the rules described in the section on dealing with missing values and can be dealt with accordingly.
In the following Python snippet we use the Vary() signature that allows varying multiple columns simultaneously or "in lockstep":
The expression returns an RVec of two RVecs: each inner vector contains the varied values for one column. The inner vectors follow the same ordering as the column names that are passed as the first argument. Besides the variation tags, in this case we also have to explicitly pass the variation name (here: "ptAndEta") as the default column name does not exist.
The above call will produce variations "ptAndEta:down" and "ptAndEta:up".
Even if a result depends on multiple variations, only one variation is applied at a time, i.e. there will be no result produced by applying multiple systematic variations at the same time. For example, in the following example snippet, the RResultMap instance all_h will contain keys "nominal", "pt:down", "pt:up", "eta:0", "eta:1", but no "pt:up&&eta:0" or similar:
Note how we passed the integer 2 instead of a list of variation tags to the second Vary() invocation: this is a shorthand that automatically generates tags 0 to N-1 (in this case 0 and 1).
RDataFrame variables/nodes are relatively cheap to copy and it's possible to both pass them to (or move them into) functions and to return them from functions. However, in general each dataframe node will have a different C++ type, which includes all available compile-time information about what that node does. One way to cope with this complication is to use template functions and/or C++14 auto return types:
A possibly simpler, C++11-compatible alternative is to take advantage of the fact that any dataframe node can be converted (implicitly or via an explicit cast) to the common type ROOT::RDF::RNode:
The conversion to ROOT::RDF::RNode is cheap, but it will introduce an extra virtual call during the RDataFrame event loop (in most cases, the resulting performance impact should be negligible). Python users can perform the conversion with the helper function ROOT.RDF.AsRNode.
ROOT::RDF::RNode also makes it simple to store RDataFrame nodes in collections, e.g. a std::vector<RNode> or a std::map<std::string, RNode>:
It's possible to schedule execution of arbitrary functions (callbacks) during the event loop. Callbacks can be used e.g. to inspect partial results of the analysis while the event loop is running, drawing a partially-filled histogram every time a certain number of new entries is processed, or displaying a progress bar while the event loop runs.
For example one can draw an up-to-date version of a result histogram every 100 entries like this:
Callbacks are registered to a ROOT::RDF::RResultPtr and must be callables that takes a reference to the result type as argument and return nothing. RDataFrame will invoke registered callbacks passing partial action results as arguments to them (e.g. a histogram filled with a part of the selected events).
Read more on ROOT::RDF::RResultPtr::OnPartialResult() and ROOT::RDF::RResultPtr::OnPartialResultSlot().
When constructing an RDataFrame object, it is possible to specify a default column list for your analysis, in the usual form of a list of strings representing branch/column names. The default column list will be used as a fallback whenever a list specific to the transformation/action is not present. RDataFrame will take as many of these columns as needed, ignoring trailing extra names if present.
Every instance of RDataFrame is created with two special columns called rdfentry_ and rdfslot_. The rdfentry_ column is of type ULong64_t and it holds the current entry number while rdfslot_ is an unsigned int holding the index of the current data processing slot. For backwards compatibility reasons, the names tdfentry_ and tdfslot_ are also accepted. These columns are ignored by operations such as Cache or Snapshot.
C++ is a statically typed language: all types must be known at compile-time. This includes the types of the TTree branches we want to work on. For filters, defined columns and some of the actions, column types are deduced from the signature of the relevant filter function/temporary column expression/action function:
If we specify an incorrect type for one of the columns, an exception with an informative message will be thrown at runtime, when the column value is actually read from the dataset: RDataFrame detects type mismatches. The same would happen if we swapped the order of "b1" and "b2" in the column list passed to Filter().
Certain actions, on the other hand, do not take a function as argument (e.g. Histo1D()), so we cannot deduce the type of the column at compile-time. In this case RDataFrame infers the type of the column from the TTree itself. This is why we never needed to specify the column types for all actions in the above snippets.
When the column type is not a common one such as int, double, char or float it is nonetheless good practice to specify it as a template parameter to the action itself, like this:
Deducing types at runtime requires the just-in-time compilation of the relevant actions, which has a small runtime overhead, so specifying the type of the columns as template parameters to the action is good practice when performance is a goal.
When strings are passed as expressions to Filter() or Define(), fundamental types are passed as constants. This avoids certaincommon mistakes such as typing x = 0 rather than x == 0:
RDataFrame strives to offer a comprehensive set of standard actions that can be performed on each event. At the same time, it allows users to inject their own action code to perform arbitrarily complex data reductions.
Through the Book() method, users can implement a custom action and have access to the same features that built-in RDataFrame actions have, e.g. hooks to events related to the start, end and execution of the event loop, or the possibility to return a lazy RResultPtr to an arbitrary type of result:
See the Book() method for more information and this tutorial for a more complete example.
Foreach() takes a callable (lambda expression, free function, functor...) and a list of columns and executes the callable on the values of those columns for each event that passes all upstream selections. It can be used to perform actions that are not already available in the interface. For example, the following snippet evaluates the root mean square of column "x":
In multi-thread runs, users are responsible for the thread-safety of the expression passed to Foreach(): thread will execute the expression concurrently. The code above would need to employ some resource protection mechanism to ensure non-concurrent writing of rms; but this is probably too much head-scratch for such a simple operation.
ForeachSlot() can help in this situation. It is an alternative version of Foreach() for which the function takes an additional "processing slot" parameter besides the columns it should be applied to. RDataFrame guarantees that ForeachSlot() will invoke the user expression with different slot parameters for different concurrent executions (see Special helper columns: rdfentry_ and rdfslot_ for more information on the slot parameter). We can take advantage of ForeachSlot() to evaluate a thread-safe root mean square of column "x":
Notice how we created one double variable for each processing slot and later merged their results via std::accumulate.
Vertically concatenating multiple trees that have the same columns (creating a logical dataset with the same columns and more rows) is trivial in RDataFrame: just pass the tree name and a list of file names to RDataFrame's constructor, or create a TChain out of the desired trees and pass that to RDataFrame.
Horizontal concatenations of trees or chains (creating a logical dataset with the same number of rows and the union of the columns of multiple trees) leverages TTree's "friend" mechanism.
Simple joins of trees that do not have the same number of rows are also possible with indexed friend trees (see below).
To use friend trees in RDataFrame, set up trees with the appropriate relationships and then instantiate an RDataFrame with the main tree:
The same applies for TChains. Columns coming from the friend trees can be referred to by their full name, like in the example above, or the friend tree name can be omitted in case the column name is not ambiguous (e.g. "MyCol" could be used instead of "myFriend.MyCol" in the example above if there is no column "MyCol" in the main tree).
Indexed friend trees provide a way to perform simple joins of multiple trees over a common column. When a certain entry in the main tree (or chain) is loaded, the friend trees (or chains) will then load an entry where the "index" columns have a value identical to the one in the main one. For example, in Python:
RDataFrame supports indexed friend TTrees from ROOT v6.24 in single-thread mode and from v6.28/02 in multi-thread mode.
RDataFrame can be interfaced with RDataSources. The ROOT::RDF::RDataSource interface defines an API that RDataFrame can use to read arbitrary columnar data formats.
RDataFrame calls into concrete RDataSource implementations to retrieve information about the data, retrieve (thread-local) readers or "cursors" for selected columns and to advance the readers to the desired data entry. Some predefined RDataSources are natively provided by ROOT such as the ROOT::RDF::RCsvDS which allows to read comma separated files:
See also FromNumpy (Python-only), FromRNTuple(), FromArrow(), FromSqlite().
As we saw, transformed dataframes can be stored as variables and reused multiple times to create modified versions of the dataset. This implicitly defines a computation graph in which several paths of filtering/creation of columns are executed simultaneously, and finally aggregated results are produced.
RDataFrame detects when several actions use the same filter or the same defined column, and only evaluates each filter or defined column once per event, regardless of how many times that result is used down the computation graph. Objects read from each column are built once and never copied, for maximum efficiency. When "upstream" filters are not passed, subsequent filters, temporary column expressions and actions are not evaluated, so it might be advisable to put the strictest filters first in the graph.
It is possible to print the computation graph from any node to obtain a DOT (graphviz) representation either on the standard output or in a file.
Invoking the function ROOT::RDF::SaveGraph() on any node that is not the head node, the computation graph of the branch the node belongs to is printed. By using the head node, the entire computation graph is printed.
Following there is an example of usage:
The generated graph can be rendered using one of the graphviz filters, e.g. dot. For instance, the image below can be generated with the following command:

RDataFrame has experimental support for verbose logging of the event loop runtimes and other interesting related information. It is activated as follows:
or in Python:
More information (e.g. start and end of each multi-thread task) is printed using ELogLevel.kDebug and even more (e.g. a full dump of the generated code that RDataFrame just-in-time-compiles) using ELogLevel.kDebug+10.
RDataFrame can be created using a dataset specification JSON file:
The input dataset specification JSON file needs to be provided by the user and it describes all necessary samples and their associated metadata information. The main required key is the "samples" (at least one sample is needed) and the required sub-keys for each sample are "trees" and "files". Additionally, one can specify a metadata dictionary for each sample in the "metadata" key.
A simple example for the formatting of the specification in the JSON file is the following:
The metadata information from the specification file can be then accessed using the DefinePerSample function. For example, to access luminosity information (stored as a double):
or sample_category information (stored as a string):
or directly the filename:
An example implementation of the "FromSpec" method is available in tutorial: df106_HiggstoFourLeptons.py, which also provides a corresponding exemplary JSON file for the dataset specification.
A progress bar showing the processed event statistics can be added to any RDataFrame program. The event statistics include elapsed time, currently processed file, currently processed events, the rate of event processing and an estimated remaining time (per file being processed). It is recorded and printed in the terminal every m events and every n seconds (by default m = 1000 and n = 1). The ProgressBar can be also added when the multithread (MT) mode is enabled.
ProgressBar is added after creating the dataframe object (df):
Alternatively, RDataFrame can be cast to an RNode first, giving the user more flexibility For example, it can be called at any computational node, such as Filter or Define, not only the head node, with no change to the ProgressBar function itself (please see the Python interface section for appropriate usage in Python):
Examples of implemented progress bars can be seen by running Higgs to Four Lepton tutorial and Dimuon tutorial.
In certain situations a dataset might be missing one or more values at one or more of its entries. For example:
For example, suppose that column "y" does not have a value for entry 42:
If the RDataFrame application reads that column, for example if a Take() action was requested, the default behaviour is to throw an exception indicating that that column is missing an entry.
The following paragraphs discuss the functionalities provided by RDataFrame to work with missing values in the dataset.
FilterAvailable and FilterMissing are specialized RDataFrame Filter operations. They take as input argument the name of a column of the dataset to watch for missing values. Like Filter, they will either keep or discard an entire entry based on whether a condition returns true or false. Specifically:
DefaultValueFor creates a node of the computation graph which just forwards the values of the columns necessary for other downstream nodes, when they are available. In case a value of the input column passed to this function is not available, the node will provide the default value passed to this function call instead. Example:
All the operations presented above only act on the particular branch of the computation graph where they are called, so that different results can be obtained by mixing and matching the filtering or providing a default value strategies:
Note that working with missing values is currently supported with a TTree-based data source. Support of this functionality for other data sources may come in the future.
RDataFrame does not treat NaNs or infinities beyond what the floating-point standards require, i.e. they will propagate to the final result. Non-finite numbers can be suppressed using Filter(), e.g.:
| TTree::Draw | ROOT::RDataFrame |
ROOT::RDataFrame df("myTree", file);
df.Filter("y > 2").Histo1D("x")->Draw();
| |
// Draw a histogram of "jet_eta" with the desired weight
tree->Draw("jet_eta", "weight*(event == 1)");
| df.Filter("event == 1").Histo1D("jet_eta", "weight")->Draw();
|
// Draw a histogram filled with values resulting from calling a method of the class of the `event` branch in the TTree.
tree->Draw("event.GetNtrack()");
| df.Define("NTrack","event.GetNtrack()").Histo1D("NTrack")->Draw();
|
// Draw only every 10th event
tree->Draw("fNtrack","fEvtHdr.fEvtNum%10 == 0");
| // Use the Filter operation together with the special RDF column: `rdfentry_`
df.Filter("rdfentry_ % 10 == 0").Histo1D("fNtrack")->Draw();
|
// object selection: for each event, fill histogram with array of selected pts
tree->Draw('Muon_pt', 'Muon_pt > 100');
| // with RDF, arrays are read as ROOT::VecOps::RVec objects
df.Define("good_pt", "Muon_pt[Muon_pt > 100]").Histo1D("good_pt")->Draw();
|
TTree::Scan()
ROOT::RDataFrame
Definition at line 50 of file RDataFrame.hxx.
Public Types | |
| using | ColumnNames_t = ROOT::RDF::ColumnNames_t |
Public Member Functions | |
| RDataFrame (ROOT::RDF::Experimental::RDatasetSpec spec) | |
| Build dataframe from an RDatasetSpec object. | |
| RDataFrame (std::string_view treeName, ::TDirectory *dirPtr, const ColumnNames_t &defaultColumns={}) | |
| RDataFrame (std::string_view treename, const std::vector< std::string > &filenames, const ColumnNames_t &defaultColumns={}) | |
| Build the dataframe. | |
| RDataFrame (std::string_view treename, std::initializer_list< std::string > filenames, const ColumnNames_t &defaultColumns={}) | |
| RDataFrame (std::string_view treeName, std::string_view filenameglob, const ColumnNames_t &defaultColumns={}) | |
| Build the dataframe. | |
| RDataFrame (std::unique_ptr< ROOT::RDF::RDataSource >, const ColumnNames_t &defaultColumns={}) | |
| Build dataframe associated to data source. | |
| RDataFrame (TTree &tree, const ColumnNames_t &defaultColumns={}) | |
| Build the dataframe. | |
| RDataFrame (ULong64_t numEntries) | |
| Build a dataframe that generates numEntries entries. | |
| RDFDescription | Describe () |
| Return information about the dataframe. | |
| ColumnNames_t | GetColumnNames () |
| Returns the names of the available columns. | |
| std::string | GetColumnType (std::string_view column) |
| Return the type of a given column as a string. | |
| ColumnNames_t | GetDatasetTopLevelFieldNames () |
| Retrieve the names of top-level field names. | |
| ColumnNames_t | GetDefinedColumnNames () |
| Returns the names of the defined columns. | |
| unsigned int | GetNFiles () |
| unsigned int | GetNRuns () const |
| Gets the number of event loops run. | |
| unsigned int | GetNSlots () const |
| Gets the number of data processing slots. | |
| RVariationsDescription | GetVariations () const |
| Return a descriptor for the systematic variations registered in this branch of the computation graph. | |
| bool | HasColumn (std::string_view columnName) |
| Checks if a column is present in the dataset. | |
Transformation | |
| operator RNode () const | |
| Cast any RDataFrame node to a common type ROOT::RDF::RNode. | |
| RInterface< RDFDetail::RFilter< F, RDFDetail::RLoopManager > > | Filter (F f, const ColumnNames_t &columns={}, std::string_view name="") |
| Append a filter to the call graph. | |
| RInterface< RDFDetail::RFilterWithMissingValues< RDFDetail::RLoopManager > > | FilterAvailable (std::string_view column) |
| Discard entries with missing values. | |
| RInterface< RDFDetail::RFilterWithMissingValues< RDFDetail::RLoopManager > > | FilterMissing (std::string_view column) |
| Keep only the entries that have missing values. | |
| RInterface< RDFDetail::RLoopManager > | Define (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Define a new column. | |
| RInterface< RDFDetail::RLoopManager > | DefineSlot (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Define a new column with a value dependent on the processing slot. | |
| RInterface< RDFDetail::RLoopManager > | DefineSlotEntry (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Define a new column with a value dependent on the processing slot and the current entry. | |
| RInterface< RDFDetail::RLoopManager > | Redefine (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Overwrite the value and/or type of an existing column. | |
| RInterface< RDFDetail::RLoopManager > | RedefineSlot (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Overwrite the value and/or type of an existing column. | |
| RInterface< RDFDetail::RLoopManager > | RedefineSlotEntry (std::string_view name, F expression, const ColumnNames_t &columns={}) |
| Overwrite the value and/or type of an existing column. | |
| RInterface< RDFDetail::RLoopManager > | DefaultValueFor (std::string_view column, const T &defaultValue) |
| In case the value in the given column is missing, provide a default value. | |
| RInterface< RDFDetail::RLoopManager > | DefinePerSample (std::string_view name, F expression) |
| Define a new column that is updated when the input sample changes. | |
| RInterface< RDFDetail::RLoopManager > | Vary (std::string_view colName, F &&expression, const ColumnNames_t &inputColumns, const std::vector< std::string > &variationTags, std::string_view variationName="") |
| Register systematic variations for a single existing column using custom variation tags. | |
| RInterface< RDFDetail::RLoopManager > | Alias (std::string_view alias, std::string_view columnName) |
| Allow to refer to a column with a different name. | |
| RResultPtr< RInterface< RLoopManager > > | Snapshot (std::string_view treename, std::string_view filename, const ColumnNames_t &columnList, const RSnapshotOptions &options=RSnapshotOptions()) |
| RInterface< RLoopManager > | Cache (const ColumnNames_t &columnList) |
| Save selected columns in memory. | |
| RInterface< RDFDetail::RRange< RDFDetail::RLoopManager > > | Range (unsigned int begin, unsigned int end, unsigned int stride=1) |
| Creates a node that filters entries based on range: [begin, end). | |
| void | Foreach (F f, const ColumnNames_t &columns={}) |
| Execute a user-defined function on each entry (instant action). | |
| void | ForeachSlot (F f, const ColumnNames_t &columns={}) |
| Execute a user-defined function requiring a processing slot index on each entry (instant action). | |
| RResultPtr< T > | Reduce (F f, std::string_view columnName="") |
| Execute a user-defined reduce operation on the values of a column. | |
| RResultPtr< ULong64_t > | Count () |
| Return the number of entries processed (lazy action). | |
| RResultPtr< COLL > | Take (std::string_view column="") |
| Return a collection of values of a column (lazy action, returns a std::vector by default). | |
| RResultPtr<::TH1D > | Histo1D (const TH1DModel &model={"", "", 128u, 0., 0.}, std::string_view vName="") |
| Fill and return a one-dimensional histogram with the values of a column (lazy action). | |
| RResultPtr<::TH2D > | Histo2D (const TH2DModel &model, std::string_view v1Name="", std::string_view v2Name="") |
| Fill and return a two-dimensional histogram (lazy action). | |
| RResultPtr<::TH3D > | Histo3D (const TH3DModel &model, std::string_view v1Name="", std::string_view v2Name="", std::string_view v3Name="") |
| Fill and return a three-dimensional histogram (lazy action). | |
| RResultPtr<::THnD > | HistoND (const THnDModel &model, const ColumnNames_t &columnList, std::string_view wName="") |
| Fill and return an N-dimensional histogram (lazy action). | |
| RResultPtr<::THnSparseD > | HistoNSparseD (const THnSparseDModel &model, const ColumnNames_t &columnList, std::string_view wName="") |
| Fill and return a sparse N-dimensional histogram (lazy action). | |
| RResultPtr< ROOT::Experimental::RHist< BinContentType > > | Hist (std::uint64_t nNormalBins, std::pair< double, double > interval, std::string_view vName) |
| Fill and return a one-dimensional RHist (lazy action). | |
| RResultPtr<::TGraph > | Graph (std::string_view x="", std::string_view y="") |
| Fill and return a TGraph object (lazy action). | |
| RResultPtr<::TGraphAsymmErrors > | GraphAsymmErrors (std::string_view x="", std::string_view y="", std::string_view exl="", std::string_view exh="", std::string_view eyl="", std::string_view eyh="") |
| Fill and return a TGraphAsymmErrors object (lazy action). | |
| RResultPtr<::TProfile > | Profile1D (const TProfile1DModel &model, std::string_view v1Name="", std::string_view v2Name="") |
| Fill and return a one-dimensional profile (lazy action). | |
| RResultPtr<::TProfile2D > | Profile2D (const TProfile2DModel &model, std::string_view v1Name="", std::string_view v2Name="", std::string_view v3Name="") |
| Fill and return a two-dimensional profile (lazy action). | |
| RResultPtr< std::decay_t< T > > | Fill (T &&model, const ColumnNames_t &columnList) |
| Return an object of type T on which T::Fill will be called once per event (lazy action). | |
| RResultPtr< TStatistic > | Stats (std::string_view value="") |
| Return a TStatistic object, filled once per event (lazy action). | |
| RResultPtr< RDFDetail::MinReturnType_t< T > > | Min (std::string_view columnName="") |
| Return the minimum of processed column values (lazy action). | |
| RResultPtr< RDFDetail::MaxReturnType_t< T > > | Max (std::string_view columnName="") |
| Return the maximum of processed column values (lazy action). | |
| RResultPtr< double > | Mean (std::string_view columnName="") |
| Return the mean of processed column values (lazy action). | |
| RResultPtr< double > | StdDev (std::string_view columnName="") |
| Return the unbiased standard deviation of processed column values (lazy action). | |
| RResultPtr< RDFDetail::SumReturnType_t< T > > | Sum (std::string_view columnName="", const RDFDetail::SumReturnType_t< T > &initValue=RDFDetail::SumReturnType_t< T >{}) |
| Return the sum of processed column values (lazy action). | |
| RResultPtr< RCutFlowReport > | Report () |
| Gather filtering statistics. | |
| std::vector< std::string > | GetFilterNames () |
| Returns the names of the filters created. | |
| RResultPtr< U > | Aggregate (AccFun aggregator, MergeFun merger, std::string_view columnName, const U &aggIdentity) |
| Execute a user-defined accumulation operation on the processed column values in each processing slot. | |
| RResultPtr< typename std::decay_t< Helper >::Result_t > | Book (Helper &&helper, const ColumnNames_t &columns={}) |
| Book execution of a custom action using a user-defined helper object. | |
| RResultPtr< RDisplay > | Display (const ColumnNames_t &columnList, size_t nRows=5, size_t nMaxCollectionElements=10) |
| Provides a representation of the columns in the dataset. | |
Protected Member Functions | |
| void | AddDefaultColumns () |
| template<typename... ColumnTypes> | |
| void | CheckAndFillDSColumns (ColumnNames_t validCols, TTraits::TypeList< ColumnTypes... > typeList) |
| void | CheckAndFillDSColumns (const std::vector< std::string > &colNames, const std::vector< const std::type_info * > &colTypeIDs) |
| void | CheckIMTDisabled (std::string_view callerName) |
| template<typename ActionTag, typename... ColTypes, typename ActionResultType, typename RDFNode, typename HelperArgType = ActionResultType, std::enable_if_t< RDFInternal::RNeedJitting< ColTypes... >::value, int > = 0> | |
| RResultPtr< ActionResultType > | CreateAction (const ColumnNames_t &columns, const std::shared_ptr< ActionResultType > &r, const std::shared_ptr< HelperArgType > &helperArg, const std::shared_ptr< RDFNode > &proxiedPtr, const int nColumns=-1, const bool vector2RVec=true) |
| Create RAction object, return RResultPtr for the action Overload for the case in which one or more column types were not specified (RTTI + jitting). | |
| template<typename ActionTag, typename... ColTypes, typename ActionResultType, typename RDFNode, typename HelperArgType = ActionResultType, std::enable_if_t<!RDFInternal::RNeedJitting< ColTypes... >::value, int > = 0> | |
| RResultPtr< ActionResultType > | CreateAction (const ColumnNames_t &columns, const std::shared_ptr< ActionResultType > &r, const std::shared_ptr< HelperArgType > &helperArg, const std::shared_ptr< RDFNode > &proxiedPtr, const int=-1) |
| Create RAction object, return RResultPtr for the action Overload for the case in which all column types were specified (no jitting). | |
| std::string | DescribeDataset () const |
| ColumnNames_t | GetColumnTypeNamesList (const ColumnNames_t &columnList) |
| RDataSource * | GetDataSource () const |
| RDFDetail::RLoopManager * | GetLoopManager () const |
| const std::shared_ptr< RDFDetail::RLoopManager > & | GetProxiedPtr () const |
| ColumnNames_t | GetValidatedColumnNames (const unsigned int nColumns, const ColumnNames_t &columns) |
| template<typename RetType> | |
| void | SanityChecksForVary (const std::vector< std::string > &colNames, const std::vector< std::string > &variationTags, std::string_view variationName) |
Protected Attributes | |
| RDFInternal::RColumnRegister | fColRegister |
| Contains the columns defined up to this node. | |
| std::shared_ptr< ROOT::Detail::RDF::RLoopManager > | fLoopManager |
| < The RLoopManager at the root of this computation graph. Never null. | |
Private Types | |
| using | RFilterBase |
| using | RLoopManager |
| using | RRangeBase |
Private Member Functions | |
| RInterface< RLoopManager > | CacheImpl (const ColumnNames_t &columnList, std::index_sequence< S... >) |
| Implementation of cache. | |
| auto | CallCreateActionWithoutColsIfPossible (const std::shared_ptr< ActionResultType > &resPtr, const std::shared_ptr< Helper > &hPtr, TTraits::TypeList< RDFDetail::RInferredType >) -> decltype(hPtr->Exec(0u), RResultPtr< ActionResultType >{}) |
| std::enable_if_t< std::is_default_constructible< RetType >::value, RInterface< RDFDetail::RLoopManager > > | DefineImpl (std::string_view name, F &&expression, const ColumnNames_t &columns, const std::string &where) |
| RInterface< RDFDetail::RLoopManager > | JittedVaryImpl (const std::vector< std::string > &colNames, std::string_view expression, const std::vector< std::string > &variationTags, std::string_view variationName, bool isSingleColumn) |
| RInterface< RDFDetail::RLoopManager > | VaryImpl (const std::vector< std::string > &colNames, F &&expression, const ColumnNames_t &inputColumns, const std::vector< std::string > &variationTags, std::string_view variationName) |
Private Attributes | |
| std::shared_ptr< RDFDetail::RLoopManager > | fProxiedPtr |
| Smart pointer to the graph node encapsulated by this RInterface. | |
#include <ROOT/RDataFrame.hxx>
Definition at line 52 of file RDataFrame.hxx.
|
privateinherited |
Definition at line 125 of file RInterface.hxx.
|
privateinherited |
Definition at line 127 of file RInterface.hxx.
|
privateinherited |
Definition at line 126 of file RInterface.hxx.
| ROOT::RDataFrame::RDataFrame | ( | std::string_view | treeName, |
| std::string_view | fileNameGlob, | ||
| const ColumnNames_t & | defaultColumns = {} ) |
Build the dataframe.
| [in] | treeName | Name of the tree contained in the directory |
| [in] | fileNameGlob | TDirectory where the tree is stored, e.g. a TFile. |
| [in] | defaultColumns | Collection of default columns. |
The filename glob supports the same type of expressions as TChain::Add(), and it is passed as-is to TChain's constructor.
The default columns are looked at in case no column is specified in the booking of actions or transformations.
Definition at line 2191 of file RDataFrame.cxx.
| ROOT::RDataFrame::RDataFrame | ( | std::string_view | datasetName, |
| const std::vector< std::string > & | fileNameGlobs, | ||
| const ColumnNames_t & | defaultColumns = {} ) |
Build the dataframe.
| [in] | datasetName | Name of the dataset contained in the directory |
| [in] | fileNameGlobs | Collection of file names of filename globs |
| [in] | defaultColumns | Collection of default columns. |
The filename globs support the same type of expressions as TChain::Add(), and each glob is passed as-is to TChain's constructor.
The default columns are looked at in case no column is specified in the booking of actions or transformations.
Definition at line 2207 of file RDataFrame.cxx.
|
inline |
Definition at line 56 of file RDataFrame.hxx.
| ROOT::RDataFrame::RDataFrame | ( | std::string_view | treeName, |
| ::TDirectory * | dirPtr, | ||
| const ColumnNames_t & | defaultColumns = {} ) |
| ROOT::RDataFrame::RDataFrame | ( | TTree & | tree, |
| const ColumnNames_t & | defaultColumns = {} ) |
Build the dataframe.
| [in] | tree | The tree or chain to be studied. |
| [in] | defaultColumns | Collection of default column names to fall back to when none is specified. |
The default columns are looked at in case no column is specified in the booking of actions or transformations.
Definition at line 2221 of file RDataFrame.cxx.
| ROOT::RDataFrame::RDataFrame | ( | ULong64_t | numEntries | ) |
Build a dataframe that generates numEntries entries.
| [in] | numEntries | The number of entries to generate. |
An empty-source dataframe constructed with a number of entries will generate those entries on the fly when some action is triggered, and it will do so for all the previously-defined columns.
Definition at line 2234 of file RDataFrame.cxx.
| ROOT::RDataFrame::RDataFrame | ( | std::unique_ptr< ROOT::RDF::RDataSource > | ds, |
| const ColumnNames_t & | defaultColumns = {} ) |
Build dataframe associated to data source.
| [in] | ds | The data source object. |
| [in] | defaultColumns | Collection of default column names to fall back to when none is specified. |
A dataframe associated to a data source will query it to access column values.
Definition at line 2247 of file RDataFrame.cxx.
| ROOT::RDataFrame::RDataFrame | ( | ROOT::RDF::Experimental::RDatasetSpec | spec | ) |
Build dataframe from an RDatasetSpec object.
| [in] | spec | The dataset specification object. |
A dataset specification includes trees and file names, as well as an optional friend list and/or entry range.
See also ROOT::RDataFrame::FromSpec().
Definition at line 2271 of file RDataFrame.cxx.
|
protectedinherited |
Definition at line 360 of file RInterfaceBase.cxx.
|
inlineinherited |
Execute a user-defined accumulation operation on the processed column values in each processing slot.
| F | The type of the aggregator callable. Automatically deduced. |
| U | The type of the aggregator variable. Must be default-constructible, copy-constructible and copy-assignable. Automatically deduced. |
| T | The type of the column to apply the reduction to. Automatically deduced. |
| [in] | aggregator | A callable with signature U(U,T) or void(U&,T), where T is the type of the column, U is the type of the aggregator variable |
| [in] | merger | A callable with signature U(U,U) or void(std::vector<U>&) used to merge the results of the accumulations of each thread |
| [in] | columnName | The column to be aggregated. If omitted, the first default column is used instead. |
| [in] | aggIdentity | The aggregator variable of each thread is initialized to this value (or is default-constructed if the parameter is omitted) |
An aggregator callable takes two values, an aggregator variable and a column value. The aggregator variable is initialized to aggIdentity or default-constructed if aggIdentity is omitted. This action calls the aggregator callable for each processed entry, passing in the aggregator variable and the value of the column columnName. If the signature is U(U,T) the aggregator variable is then copy-assigned the result of the execution of the callable. Otherwise the signature of aggregator must be void(U&,T).
The merger callable is used to merge the partial accumulation results of each processing thread. It is only called in multi-thread executions. If its signature is U(U,U) the aggregator variables of each thread are merged two by two. If its signature is void(std::vector<U>& a) it is assumed that it merges all aggregators in a[0].
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Example usage:
Definition at line 3518 of file RInterface.hxx.
|
inlineinherited |
Allow to refer to a column with a different name.
| [in] | alias | name of the column alias |
| [in] | columnName | of the column to be aliased |
Aliasing an alias is supported.
Definition at line 1293 of file RInterface.hxx.
|
inlineinherited |
Book execution of a custom action using a user-defined helper object.
| FirstColumn | The type of the first column used by this action. Inferred together with OtherColumns if not present. |
| OtherColumns | A list of the types of the other columns used by this action |
| Helper | The type of the user-defined helper. See below for the required interface it should expose. |
| [in] | helper | The Action Helper to be scheduled. |
| [in] | columns | The names of the columns on which the helper acts. |
This method books a custom action for execution. The behavior of the action is completely dependent on the Helper object provided by the caller. The required interface for the helper is described below (more methods that the ones required can be present, e.g. a constructor that takes the number of worker threads is usually useful):
If these methods are implemented they enable extra functionality as per the description below.
In case Book is called without specifying column types as template arguments, corresponding typed code will be just-in-time compiled by RDataFrame. In that case the Helper class needs to be known to the ROOT interpreter.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
See this tutorial for an example implementation of an action helper.
It is also possible to inspect the code used by built-in RDataFrame actions at ActionHelpers.hxx.
Definition at line 3624 of file RInterface.hxx.
|
inlineinherited |
Save selected columns in memory.
| ColumnTypes | variadic list of branch/column types. |
| [in] | columnList | columns to be cached in memory. |
This action returns a new RDataFrame object, completely detached from the originating RDataFrame. The new dataframe only contains the cached columns and stores their content in memory for fast, zero-copy subsequent access.
Use Cache if you know you will only need a subset of the (Filtered) data that fits in memory and that will be accessed many times.
Types and columns specified:
Types inferred and columns specified (this invocation relies on jitting):
Types inferred and columns selected with a regexp (this invocation relies on jitting):
Definition at line 1637 of file RInterface.hxx.
|
inlineprivateinherited |
Implementation of cache.
Definition at line 3803 of file RInterface.hxx.
|
inlineprivateinherited |
Definition at line 3895 of file RInterface.hxx.
|
inlineprotectedinherited |
Definition at line 142 of file RInterfaceBase.hxx.
|
inlineprotectedinherited |
Definition at line 148 of file RInterfaceBase.hxx.
|
protectedinherited |
Definition at line 351 of file RInterfaceBase.cxx.
|
inlineinherited |
Return the number of entries processed (lazy action).
Useful e.g. for counting the number of entries passing a certain filter (see also Report). This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 1903 of file RInterface.hxx.
|
inlineprotectedinherited |
Create RAction object, return RResultPtr for the action Overload for the case in which one or more column types were not specified (RTTI + jitting).
This overload has a nColumns optional argument. If present, the number of required columns for this action is taken equal to nColumns, otherwise it is assumed to be sizeof...(ColTypes).
Definition at line 187 of file RInterfaceBase.hxx.
|
inlineprotectedinherited |
Create RAction object, return RResultPtr for the action Overload for the case in which all column types were specified (no jitting).
For most actions, r and helperArg will refer to the same object, because the only argument to forward to the action helper is the result value itself. We need the distinction for actions such as Snapshot or Cache, for which the constructor arguments of the action helper are different from the returned value.
Definition at line 163 of file RInterfaceBase.hxx.
|
inlineinherited |
In case the value in the given column is missing, provide a default value.
| T | The type of the column |
| [in] | column | Column name where missing values should be replaced by the given default value |
| [in] | defaultValue | Value to provide instead of a missing value |
This operation is useful in case an entry of the dataset is incomplete, i.e. if one or more of the columns do not have valid values. It does not modify the values of the column, but in case any entry is missing, it will provide the default value to downstream nodes instead.
Use cases include:
Definition at line 682 of file RInterface.hxx.
|
inlineinherited |
Define a new column.
| [in] | name | The name of the defined column. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. This callable must be thread safe when used with multiple threads. |
| [in] | columns | Names of the columns/branches in input to the producer function. |
Define a column that will be visible from all subsequent nodes of the functional chain. The expression is only evaluated for entries that pass all the preceding filters. A new variable is created called name, accessible as if it was contained in the dataset from subsequent transformations/actions.
Use cases include:
An exception is thrown if the name of the new column is already in use in this branch of the computation graph. Note that the callable must be thread safe when called from multiple threads. Use DefineSlot() if needed.
Definition at line 453 of file RInterface.hxx.
|
inlineprivateinherited |
Definition at line 3742 of file RInterface.hxx.
|
inlineinherited |
Define a new column that is updated when the input sample changes.
| [in] | name | The name of the defined column. |
| [in] | expression | A C++ callable that computes the new value of the defined column. |
The signature of the callable passed as second argument should be T(unsigned int slot, const ROOT::RDF::RSampleInfo &id) where:
DefinePerSample() is useful to e.g. define a quantity that depends on which TTree in which TFile is being processed or to inject a callback into the event loop that is only called when the processing of a new sample starts rather than at every entry.
The callable will be invoked once per input TTree or once per multi-thread task, whichever is more often.
Definition at line 745 of file RInterface.hxx.
|
inlineinherited |
Define a new column with a value dependent on the processing slot.
| [in] | name | The name of the defined column. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. |
| [in] | columns | Names of the columns/branches in input to the producer function (excluding the slot number). |
This alternative implementation of Define is meant as a helper to evaluate new column values in a thread-safe manner. The expression must be a callable of signature R(unsigned int, T1, T2, ...) where T1, T2... are the types of the columns that the expression takes as input. The first parameter is reserved for an unsigned integer representing a "slot number". RDataFrame guarantees that different threads will invoke the expression with different slot numbers - slot numbers will range from zero to ROOT::GetThreadPoolSize()-1. Note that there is no guarantee as to how often each slot will be reached during the event loop.
The following two calls are equivalent, although DefineSlot is slightly more performant:
See Define() for more information.
Definition at line 483 of file RInterface.hxx.
|
inlineinherited |
Define a new column with a value dependent on the processing slot and the current entry.
| [in] | name | The name of the defined column. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. |
| [in] | columns | Names of the columns/branches in input to the producer function (excluding slot and entry). |
This alternative implementation of Define is meant as a helper in writing entry-specific, thread-safe custom columns. The expression must be a callable of signature R(unsigned int, ULong64_t, T1, T2, ...) where T1, T2... are the types of the columns that the expression takes as input. The first parameter is reserved for an unsigned integer representing a "slot number". RDataFrame guarantees that different threads will invoke the expression with different slot numbers - slot numbers will range from zero to ROOT::GetThreadPoolSize()-1. Note that there is no guarantee as to how often each slot will be reached during the event loop. The second parameter is reserved for a ULong64_t representing the current entry being processed by the current thread.
The following two Defines are equivalent, although DefineSlotEntry is slightly more performant:
See Define() for more information.
Definition at line 514 of file RInterface.hxx.
|
inherited |
Return information about the dataframe.
This convenience function describes the dataframe and combines the following information:
This is not an action nor a transformation, just a query to the RDataFrame object. The result is dependent on the node from which this method is called, e.g. the list of defined columns returned by GetDefinedColumnNames().
Please note that this is a convenience feature and the layout of the output can be subject to change and should be parsed via RDFDescription methods.
Definition at line 178 of file RInterfaceBase.cxx.
|
protectedinherited |
Definition at line 35 of file RInterfaceBase.cxx.
|
inlineinherited |
Provides a representation of the columns in the dataset.
| ColumnTypes | variadic list of branch/column types. |
| [in] | columnList | Names of the columns to be displayed. |
| [in] | nRows | Number of events for each column to be displayed. |
| [in] | nMaxCollectionElements | Maximum number of collection elements to display per row. |
This function returns a RResultPtr<RDisplay> containing all the entries to be displayed, organized in a tabular form. RDisplay will either print on the standard output a summarized version through RDisplay::Print() or will return a complete version through RDisplay::AsString().
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Example usage:
Definition at line 3669 of file RInterface.hxx.
|
inlineinherited |
Return an object of type T on which T::Fill will be called once per event (lazy action).
Type T must provide at least:
| FirstColumn | The first type of the column the values of which are used to fill the object. Inferred together with OtherColumns if not present. |
| OtherColumns | A list of the other types of the columns the values of which are used to fill the object. |
| T | The type of the object to fill. Automatically deduced. |
| [in] | model | The model to be considered to build the new return value. |
| [in] | columnList | A list containing the names of the columns that will be passed when calling Fill |
The user gives up ownership of the model object. The list of column names to be used for filling must always be specified. This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3173 of file RInterface.hxx.
|
inlineinherited |
Append a filter to the call graph.
| [in] | f | Function, lambda expression, functor class or any other callable object. It must return a bool signalling whether the event has passed the selection (true) or not (false). |
| [in] | columns | Names of the columns/branches in input to the filter function. |
| [in] | name | Optional name of this filter. See Report. |
Append a filter node at the point of the call graph corresponding to the object this method is called on. The callable f should not have side-effects (e.g. modification of an external or static variable) to ensure correct results when implicit multi-threading is active.
RDataFrame only evaluates filters when necessary: if multiple filters are chained one after another, they are executed in order and the first one returning false causes the event to be discarded. Even if multiple actions or transformations depend on the same filter, it is executed once per entry. If its result is requested more than once, the cached result is served.
Definition at line 235 of file RInterface.hxx.
|
inlineinherited |
Discard entries with missing values.
| [in] | column | Column name whose entries with missing values should be discarded |
This operation is useful in case an entry of the dataset is incomplete, i.e. if one or more of the columns do not have valid values. If the value of the input column is missing for an entry, the entire entry will be discarded from the rest of this branch of the computation graph.
Use cases include:
Definition at line 348 of file RInterface.hxx.
|
inlineinherited |
Keep only the entries that have missing values.
| [in] | column | Column name whose entries with missing values should be kept |
This operation is useful in case an entry of the dataset is incomplete, i.e. if one or more of the columns do not have valid values. It only keeps the entries for which the value of the input column is missing.
Use cases include:
Definition at line 399 of file RInterface.hxx.
|
inlineinherited |
Execute a user-defined function on each entry (instant action).
| [in] | f | Function, lambda expression, functor class or any other callable object performing user defined calculations. |
| [in] | columns | Names of the columns/branches in input to the user function. |
The callable f is invoked once per entry. This is an instant action: upon invocation, an event loop as well as execution of all scheduled actions is triggered. Users are responsible for the thread-safety of this callable when executing with implicit multi-threading enabled (i.e. ROOT::EnableImplicitMT).
Definition at line 1785 of file RInterface.hxx.
|
inlineinherited |
Execute a user-defined function requiring a processing slot index on each entry (instant action).
| [in] | f | Function, lambda expression, functor class or any other callable object performing user defined calculations. |
| [in] | columns | Names of the columns/branches in input to the user function. |
Same as Foreach, but the user-defined function takes an extra unsigned int as its first parameter, the processing slot index. This slot index will be assigned a different value, 0 to poolSize - 1, for each thread of execution. This is meant as a helper in writing thread-safe Foreach actions when using RDataFrame after ROOT::EnableImplicitMT(). The user-defined processing callable is able to follow different streams of processing indexed by the first parameter. ForeachSlot works just as well with single-thread execution: in that case slot will always be 0.
Definition at line 1815 of file RInterface.hxx.
|
inherited |
Returns the names of the available columns.
This is not an action nor a transformation, just a query to the RDataFrame object.
Definition at line 77 of file RInterfaceBase.cxx.
|
inherited |
Return the type of a given column as a string.
This is not an action nor a transformation, just a query to the RDataFrame object.
Definition at line 138 of file RInterfaceBase.cxx.
|
protectedinherited |
Definition at line 341 of file RInterfaceBase.cxx.
|
inherited |
Retrieve the names of top-level field names.
For data sources that support hierarchical dataset schemas, such as TTree or RNTuple, this function will retrieve the names of top-level fields. For example, if the schema contains a user class with a data member, only the name of the top-level field containing the user class object would be reported, but not the name of the data member sub-field.
For all other data sources, returns the list of all available dataset columns.
Definition at line 113 of file RInterfaceBase.cxx.
|
inlineprotectedinherited |
Definition at line 134 of file RInterfaceBase.hxx.
|
inherited |
Returns the names of the defined columns.
This is not an action nor a transformation, just a simple utility to get the columns names that have been defined up to the node. If no column has been defined, e.g. on a root node, it returns an empty collection.
Definition at line 250 of file RInterfaceBase.cxx.
|
inlineinherited |
Returns the names of the filters created.
If called on a root node, all the filters in the computation graph will be printed. For any other node, only the filters upstream of that node. Filters without a name are printed as "Unnamed Filter" This is not an action nor a transformation, just a query to the RDataFrame object.
Definition at line 3466 of file RInterface.hxx.
|
inlineprotectedinherited |
Definition at line 133 of file RInterfaceBase.hxx.
|
inherited |
Definition at line 27 of file RInterfaceBase.cxx.
|
inherited |
Gets the number of event loops run.
This method returns the number of events loops run so far by this RDataFrame instance.
Example usage:
Definition at line 336 of file RInterfaceBase.cxx.
|
inherited |
Gets the number of data processing slots.
This method returns the number of data processing slots used by this RDataFrame instance. This number is influenced by the global switch ROOT::EnableImplicitMT().
Example usage:
Definition at line 317 of file RInterfaceBase.cxx.
|
inlineprotectedinherited |
Definition at line 3922 of file RInterface.hxx.
|
inlineprotectedinherited |
Definition at line 136 of file RInterfaceBase.hxx.
|
inherited |
Return a descriptor for the systematic variations registered in this branch of the computation graph.
This is not an action nor a transformation, just a simple utility to inspect the systematic variations that have been registered with Vary() up to this node. When called on the root node, it returns an empty descriptor.
Definition at line 275 of file RInterfaceBase.cxx.
|
inlineinherited |
Fill and return a TGraph object (lazy action).
| X | The type of the column used to fill the x axis. |
| Y | The type of the column used to fill the y axis. |
| [in] | x | The name of the column that will fill the x axis. |
| [in] | y | The name of the column that will fill the y axis. |
Columns can be of a container type (e.g. std::vector<double>), in which case the TGraph is filled with each one of the elements of the container. If Multithreading is enabled, the order in which points are inserted is undefined. If the Graph has to be drawn, it is suggested to the user to sort it on the x before printing. A name and a title to the TGraph is given based on the input column names.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 2850 of file RInterface.hxx.
|
inlineinherited |
Fill and return a TGraphAsymmErrors object (lazy action).
| [in] | x | The name of the column that will fill the x axis. |
| [in] | y | The name of the column that will fill the y axis. |
| [in] | exl | The name of the column of X low errors |
| [in] | exh | The name of the column of X high errors |
| [in] | eyl | The name of the column of Y low errors |
| [in] | eyh | The name of the column of Y high errors |
Columns can be of a container type (e.g. std::vector<double>), in which case the graph is filled with each one of the elements of the container. If Multithreading is enabled, the order in which points are inserted is undefined.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
GraphAsymmErrors should also be used for the cases in which values associated only with one of the axes have associated errors. For example, only ey exist and ex are equal to zero. In such cases, user should do the following:
Definition at line 2915 of file RInterface.hxx.
|
inherited |
Checks if a column is present in the dataset.
This method checks if a column is part of the input ROOT dataset, has been defined or can be provided by the data source.
Example usage:
Definition at line 294 of file RInterfaceBase.cxx.
|
inlineinherited |
Fill and return a one-dimensional RHist (lazy action).
| BinContentType | The bin content type of the returned RHist. |
| [in] | nNormalBins | The returned histogram will be constructed using this number of normal bins. |
| [in] | interval | The axis interval of the constructed histogram (lower end inclusive, upper end exclusive). |
| [in] | vName | The name of the column that will fill the histogram. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 2562 of file RInterface.hxx.
|
inlineinherited |
Fill and return a one-dimensional histogram with the values of a column (lazy action).
| V | The type of the column used to fill the histogram. |
| [in] | model | The returned histogram will be constructed using this as a model. |
| [in] | vName | The name of the column that will fill the histogram. |
Columns can be of a container type (e.g. std::vector<double>), in which case the histogram is filled with each one of the elements of the container. In case multiple columns of container type are provided (e.g. values and weights) they must have the same length for each one of the events (but possibly different lengths between events). This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 1978 of file RInterface.hxx.
|
inlineinherited |
Fill and return a two-dimensional histogram (lazy action).
| V1 | The type of the column used to fill the x axis of the histogram. |
| V2 | The type of the column used to fill the y axis of the histogram. |
| [in] | model | The returned histogram will be constructed using this as a model. |
| [in] | v1Name | The name of the column that will fill the x axis. |
| [in] | v2Name | The name of the column that will fill the y axis. |
Columns can be of a container type (e.g. std::vector<double>), in which case the histogram is filled with each one of the elements of the container. In case multiple columns of container type are provided (e.g. values and weights) they must have the same length for each one of the events (but possibly different lengths between events). This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 2130 of file RInterface.hxx.
|
inlineinherited |
Fill and return a three-dimensional histogram (lazy action).
| V1 | The type of the column used to fill the x axis of the histogram. Inferred if not present. |
| V2 | The type of the column used to fill the y axis of the histogram. Inferred if not present. |
| V3 | The type of the column used to fill the z axis of the histogram. Inferred if not present. |
| [in] | model | The returned histogram will be constructed using this as a model. |
| [in] | v1Name | The name of the column that will fill the x axis. |
| [in] | v2Name | The name of the column that will fill the y axis. |
| [in] | v3Name | The name of the column that will fill the z axis. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 2227 of file RInterface.hxx.
|
inlineinherited |
Fill and return an N-dimensional histogram (lazy action).
| FirstColumn | The first type of the column the values of which are used to fill the object. Inferred if not present. |
| OtherColumns | A list of the other types of the columns the values of which are used to fill the object. |
| [in] | model | The returned histogram will be constructed using this as a model. |
| [in] | columnList | A list containing the names of the columns that will be passed when calling Fill. |
| [in] | wName | The name of the column that will provide the weights. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. See RResultPtr documentation.
Definition at line 2326 of file RInterface.hxx.
|
inlineinherited |
Fill and return a sparse N-dimensional histogram (lazy action).
| FirstColumn | The first type of the column the values of which are used to fill the object. Inferred if not present. |
| OtherColumns | A list of the other types of the columns the values of which are used to fill the object. |
| [in] | model | The returned histogram will be constructed using this as a model. |
| [in] | columnList | A list containing the names of the columns that will be passed when calling Fill. |
| [in] | wName | The name of the column that will provide the weights. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. See RResultPtr documentation.
Definition at line 2447 of file RInterface.hxx.
|
inlineprivateinherited |
Definition at line 3858 of file RInterface.hxx.
|
inlineinherited |
Return the maximum of processed column values (lazy action).
| T | The type of the branch/column. |
| [in] | columnName | The name of the branch/column to be treated. |
If T is not specified, RDataFrame will infer it from the data and just-in-time compile the correct template specialization of this method. If the type of the column is inferred, the return type is double, the type of the column otherwise.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3307 of file RInterface.hxx.
|
inlineinherited |
Return the mean of processed column values (lazy action).
| T | The type of the branch/column. |
| [in] | columnName | The name of the branch/column to be treated. |
If T is not specified, RDataFrame will infer it from the data and just-in-time compile the correct template specialization of this method. Note that internally, the summations are executed with Kahan sums in double precision, irrespective of the type of column that is read.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3338 of file RInterface.hxx.
|
inlineinherited |
Return the minimum of processed column values (lazy action).
| T | The type of the branch/column. |
| [in] | columnName | The name of the branch/column to be treated. |
If T is not specified, RDataFrame will infer it from the data and just-in-time compile the correct template specialization of this method. If the type of the column is inferred, the return type is double, the type of the column otherwise.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3277 of file RInterface.hxx.
|
inlineinherited |
Cast any RDataFrame node to a common type ROOT::RDF::RNode.
Different RDataFrame methods return different C++ types. All nodes, however, can be cast to this common type at the cost of a small performance penalty. This allows, for example, storing RDataFrame nodes in a vector, or passing them around via (non-template, C++11) helper functions. Example usage:
Note that it is not a problem to pass RNode's by value.
Definition at line 190 of file RInterface.hxx.
|
inlineinherited |
Fill and return a one-dimensional profile (lazy action).
| V1 | The type of the column the values of which are used to fill the profile. Inferred if not present. |
| V2 | The type of the column the values of which are used to fill the profile. Inferred if not present. |
| [in] | model | The model to be considered to build the new return value. |
| [in] | v1Name | The name of the column that will fill the x axis. |
| [in] | v2Name | The name of the column that will fill the y axis. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 2962 of file RInterface.hxx.
|
inlineinherited |
Fill and return a two-dimensional profile (lazy action).
| V1 | The type of the column used to fill the x axis of the histogram. Inferred if not present. |
| V2 | The type of the column used to fill the y axis of the histogram. Inferred if not present. |
| V3 | The type of the column used to fill the z axis of the histogram. Inferred if not present. |
| [in] | model | The returned profile will be constructed using this as a model. |
| [in] | v1Name | The name of the column that will fill the x axis. |
| [in] | v2Name | The name of the column that will fill the y axis. |
| [in] | v3Name | The name of the column that will fill the z axis. |
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3063 of file RInterface.hxx.
|
inlineinherited |
Creates a node that filters entries based on range: [begin, end).
| [in] | begin | Initial entry number considered for this range. |
| [in] | end | Final entry number (excluded) considered for this range. 0 means that the range goes until the end of the dataset. |
| [in] | stride | Process one entry of the [begin, end) range every stride entries. Must be strictly greater than 0. |
Note that in case of previous Ranges and Filters the selected range refers to the transformed dataset. Ranges are only available if EnableImplicitMT has not been called. Multi-thread ranges are not supported.
Definition at line 1744 of file RInterface.hxx.
|
inlineinherited |
Overwrite the value and/or type of an existing column.
| [in] | name | The name of the column to redefine. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. |
| [in] | columns | Names of the columns/branches in input to the expression. |
The old value of the column can be used as an input for the expression.
An exception is thrown in case the column to redefine does not already exist. See Define() for more information.
Definition at line 572 of file RInterface.hxx.
|
inlineinherited |
Overwrite the value and/or type of an existing column.
| [in] | name | The name of the column to redefine. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. |
| [in] | columns | Names of the columns/branches in input to the producer function (excluding slot). |
The old value of the column can be used as an input for the expression. An exception is thrown in case the column to redefine does not already exist.
See DefineSlot() for more information.
Definition at line 591 of file RInterface.hxx.
|
inlineinherited |
Overwrite the value and/or type of an existing column.
| [in] | name | The name of the column to redefine. |
| [in] | expression | Function, lambda expression, functor class or any other callable object producing the defined value. Returns the value that will be assigned to the defined column. |
| [in] | columns | Names of the columns/branches in input to the producer function (excluding slot and entry). |
The old value of the column can be used as an input for the expression. An exception is thrown in case the column to re-define does not already exist.
See DefineSlotEntry() for more information.
Definition at line 610 of file RInterface.hxx.
|
inlineinherited |
Execute a user-defined reduce operation on the values of a column.
| F | The type of the reduce callable. Automatically deduced. |
| T | The type of the column to apply the reduction to. Automatically deduced. |
| [in] | f | A callable with signature T(T,T) |
| [in] | columnName | The column to be reduced. If omitted, the first default column is used instead. |
A reduction takes two values of a column and merges them into one (e.g. by summing them, taking the maximum, etc). This action performs the specified reduction operation on all processed column values, returning a single value of the same type. The callable f must satisfy the general requirements of a processing function besides having signature T(T,T) where T is the type of column columnName.
The returned reduced value of each thread (e.g. the initial value of a sum) is initialized to a default-constructed T object. This is commonly expected to be the neutral/identity element for the specific reduction operation f (e.g. 0 for a sum, 1 for a product). If a default-constructed T does not satisfy this requirement, users should explicitly specify an initialization value for T by calling the appropriate Reduce overload.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 1862 of file RInterface.hxx.
|
inlineinherited |
Gather filtering statistics.
Calling Report on the main RDataFrame object gathers stats for all named filters in the call graph. Calling this method on a stored chain state (i.e. a graph node different from the first) gathers the stats for all named filters in the chain section between the original RDataFrame and that node (included). Stats are gathered in the same order as the named filters have been added to the graph. A RResultPtr<RCutFlowReport> is returned to allow inspection of the effects cuts had.
This action is lazy: upon invocation of this method the calculation is booked but not executed. See RResultPtr documentation.
Definition at line 3431 of file RInterface.hxx.
|
inlineprotectedinherited |
Definition at line 66 of file RInterfaceBase.hxx.
|
inlineinherited |
Definition at line 1320 of file RInterface.hxx.
|
inlineinherited |
Return a TStatistic object, filled once per event (lazy action).
| V | The type of the value column |
| [in] | value | The name of the column with the values to fill the statistics with. |
Definition at line 3199 of file RInterface.hxx.
|
inlineinherited |
Return the unbiased standard deviation of processed column values (lazy action).
| T | The type of the branch/column. |
| [in] | columnName | The name of the branch/column to be treated. |
If T is not specified, RDataFrame will infer it from the data and just-in-time compile the correct template specialization of this method.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3366 of file RInterface.hxx.
|
inlineinherited |
Return the sum of processed column values (lazy action).
| T | The type of the branch/column. |
| [in] | columnName | The name of the branch/column. |
| [in] | initValue | Optional initial value for the sum. If not present, the column values must be default-constructible. |
If T is not specified, RDataFrame will infer it from the data and just-in-time compile the correct template specialization of this method. If the type of the column is inferred, the return type is double, the type of the column otherwise.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 3398 of file RInterface.hxx.
|
inlineinherited |
Return a collection of values of a column (lazy action, returns a std::vector by default).
| T | The type of the column. |
| COLL | The type of collection used to store the values. |
| [in] | column | The name of the column to collect the values of. |
The collection type to be specified for C-style array columns is RVec<T>: in this case the returned collection is a std::vector<RVec<T>>.
This action is lazy: upon invocation of this method the calculation is booked but not executed. Also see RResultPtr.
Definition at line 1935 of file RInterface.hxx.
|
inlineinherited |
Register systematic variations for a single existing column using custom variation tags.
| [in] | colName | name of the column for which varied values are provided. |
| [in] | expression | a callable that evaluates the varied values for the specified columns. The callable can take any column values as input, similarly to what happens during Filter and Define calls. It must return an RVec of varied values, one for each variation tag, in the same order as the tags. |
| [in] | inputColumns | the names of the columns to be passed to the callable. |
| [in] | variationTags | names for each of the varied values, e.g. "up" and "down". |
| [in] | variationName | a generic name for this set of varied values, e.g. "ptvariation". |
Vary provides a natural and flexible syntax to define systematic variations that automatically propagate to Filters, Defines and results. RDataFrame usage of columns with attached variations does not change, but for results that depend on any varied quantity, a map/dictionary of varied results can be produced with ROOT::RDF::Experimental::VariationsFor (see the example below).
The dictionary will contain a "nominal" value (accessed with the "nominal" key) for the unchanged result, and values for each of the systematic variations that affected the result (via upstream Filters or via direct or indirect dependencies of the column values on some registered variations). The keys will be a composition of variation names and tags, e.g. "pt:up" and "pt:down" for the example below.
In the following example we add up/down variations of pt and fill a histogram with a quantity that depends on pt. We automatically obtain three histograms in output ("nominal", "pt:up" and "pt:down"):
RDataFrame computes all variations as part of a single loop over the data. In particular, this means that I/O and computation of values shared among variations only happen once for all variations. Thus, the event loop run-time typically scales much better than linearly with the number of variations.
RDataFrame lazily computes the varied values required to produce the outputs of VariationsFor(). If VariationsFor() was not called for a result, the computations are only run for the nominal case.
See other overloads for examples when variations are added for multiple existing columns, or when the tags are auto-generated instead of being directly defined.
Definition at line 870 of file RInterface.hxx.
|
inlineprivateinherited |
Definition at line 3825 of file RInterface.hxx.
|
protectedinherited |
Contains the columns defined up to this node.
Definition at line 55 of file RInterfaceBase.hxx.
|
protectedinherited |
< The RLoopManager at the root of this computation graph. Never null.
Definition at line 52 of file RInterfaceBase.hxx.
|
privateinherited |
Smart pointer to the graph node encapsulated by this RInterface.
Definition at line 142 of file RInterface.hxx.