␛[1mRooFit v3.60 -- Developed by Wouter Verkerke and David Kirkby␛[0m
Copyright (C) 2000-2013 NIKHEF, University of California & Stanford University
All rights reserved, please read http://roofit.sourceforge.net/license.txt
RooWorkspace(combined) combined contents
variables
---------
(Lumi,SigXsecOverSM,alpha_syst1,alpha_syst2,alpha_syst3,binWidth_obs_x_channel1_0,binWidth_obs_x_channel1_1,binWidth_obs_x_channel1_2,channelCat,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1,nom_alpha_syst1,nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1,nominalLumi,obs_x_channel1,weightVar)
p.d.f.s
-------
RooGaussian::alpha_syst1Constraint[ x=alpha_syst1 mean=nom_alpha_syst1 sigma=1 ] = 1
RooGaussian::alpha_syst2Constraint[ x=alpha_syst2 mean=nom_alpha_syst2 sigma=1 ] = 1
RooGaussian::alpha_syst3Constraint[ x=alpha_syst3 mean=nom_alpha_syst3 sigma=1 ] = 1
RooRealSumPdf::channel1_model[ binWidth_obs_x_channel1_0 * L_x_signal_channel1_overallSyst_x_Exp + binWidth_obs_x_channel1_1 * L_x_background1_channel1_overallSyst_x_StatUncert + binWidth_obs_x_channel1_2 * L_x_background2_channel1_overallSyst_x_StatUncert ] = 220
RooPoisson::gamma_stat_channel1_bin_0_constraint[ x=nom_gamma_stat_channel1_bin_0 mean=gamma_stat_channel1_bin_0_poisMean ] = 0.019943
RooPoisson::gamma_stat_channel1_bin_1_constraint[ x=nom_gamma_stat_channel1_bin_1 mean=gamma_stat_channel1_bin_1_poisMean ] = 0.039861
RooGaussian::lumiConstraint[ x=Lumi mean=nominalLumi sigma=0.1 ] = 1
RooProdPdf::model_channel1[ lumiConstraint * alpha_syst1Constraint * alpha_syst2Constraint * alpha_syst3Constraint * gamma_stat_channel1_bin_0_constraint * gamma_stat_channel1_bin_1_constraint * channel1_model(obs_x_channel1) ] = 0.174888
RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.174888
functions
--------
RooProduct::L_x_background1_channel1_overallSyst_x_StatUncert[ Lumi * background1_channel1_overallSyst_x_StatUncert ] = 0
RooProduct::L_x_background2_channel1_overallSyst_x_StatUncert[ Lumi * background2_channel1_overallSyst_x_StatUncert ] = 100
RooProduct::L_x_signal_channel1_overallSyst_x_Exp[ Lumi * signal_channel1_overallSyst_x_Exp ] = 10
RooStats::HistFactory::FlexibleInterpVar::background1_channel1_epsilon[ paramList=(alpha_syst2) ] = 1
RooHistFunc::background1_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 0
RooProduct::background1_channel1_overallSyst_x_Exp[ background1_channel1_nominal * background1_channel1_epsilon ] = 0
RooProduct::background1_channel1_overallSyst_x_StatUncert[ mc_stat_channel1 * background1_channel1_overallSyst_x_Exp ] = 0
RooStats::HistFactory::FlexibleInterpVar::background2_channel1_epsilon[ paramList=(alpha_syst3) ] = 1
RooHistFunc::background2_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 100
RooProduct::background2_channel1_overallSyst_x_Exp[ background2_channel1_nominal * background2_channel1_epsilon ] = 100
RooProduct::background2_channel1_overallSyst_x_StatUncert[ mc_stat_channel1 * background2_channel1_overallSyst_x_Exp ] = 100
RooProduct::gamma_stat_channel1_bin_0_poisMean[ gamma_stat_channel1_bin_0 * gamma_stat_channel1_bin_0_tau ] = 400
RooProduct::gamma_stat_channel1_bin_1_poisMean[ gamma_stat_channel1_bin_1 * gamma_stat_channel1_bin_1_tau ] = 100
ParamHistFunc::mc_stat_channel1[ ] = 1
RooStats::HistFactory::FlexibleInterpVar::signal_channel1_epsilon[ paramList=(alpha_syst1) ] = 1
RooHistFunc::signal_channel1_nominal[ depList=(obs_x_channel1) depList=(obs_x_channel1) ] = 10
RooProduct::signal_channel1_overallNorm_x_sigma_epsilon[ SigXsecOverSM * signal_channel1_epsilon ] = 1
RooProduct::signal_channel1_overallSyst_x_Exp[ signal_channel1_nominal * signal_channel1_overallNorm_x_sigma_epsilon ] = 10
datasets
--------
RooDataSet::asimovData(obs_x_channel1,weightVar,channelCat)
RooDataSet::obsData(channelCat,obs_x_channel1)
embedded datasets (in pdfs and functions)
-----------------------------------------
RooDataHist::signal_channel1nominalDHist(obs_x_channel1)
RooDataHist::background1_channel1nominalDHist(obs_x_channel1)
RooDataHist::background2_channel1nominalDHist(obs_x_channel1)
parameter snapshots
-------------------
NominalParamValues = (nom_alpha_syst2=0[C],nom_alpha_syst3=0[C],nom_gamma_stat_channel1_bin_0=400[C],nom_gamma_stat_channel1_bin_1=100[C],weightVar=0,obs_x_channel1=1.75,Lumi=1[C],nominalLumi=1[C],alpha_syst1=0[C],nom_alpha_syst1=0[C],alpha_syst2=0,alpha_syst3=0,gamma_stat_channel1_bin_0=1,gamma_stat_channel1_bin_1=1,SigXsecOverSM=1,binWidth_obs_x_channel1_0=2[C],binWidth_obs_x_channel1_1=2[C],binWidth_obs_x_channel1_2=2[C])
named sets
----------
ModelConfig_GlobalObservables:(nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
ModelConfig_NuisParams:(alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
ModelConfig_Observables:(obs_x_channel1,weightVar,channelCat)
ModelConfig_POI:(SigXsecOverSM)
globalObservables:(nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
observables:(obs_x_channel1,weightVar,channelCat)
generic objects
---------------
RooStats::ModelConfig::ModelConfig
=== Using the following for ModelConfigB_only ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.174888
Snapshot:
1) 0x55b1b4f2b680 RooRealVar:: SigXsecOverSM = 0 L(0 - 3) "SigXsecOverSM"
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,weightVar,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.174888
Snapshot:
1) 0x55b1b4f2b680 RooRealVar:: SigXsecOverSM = 1 L(0 - 3) "SigXsecOverSM"
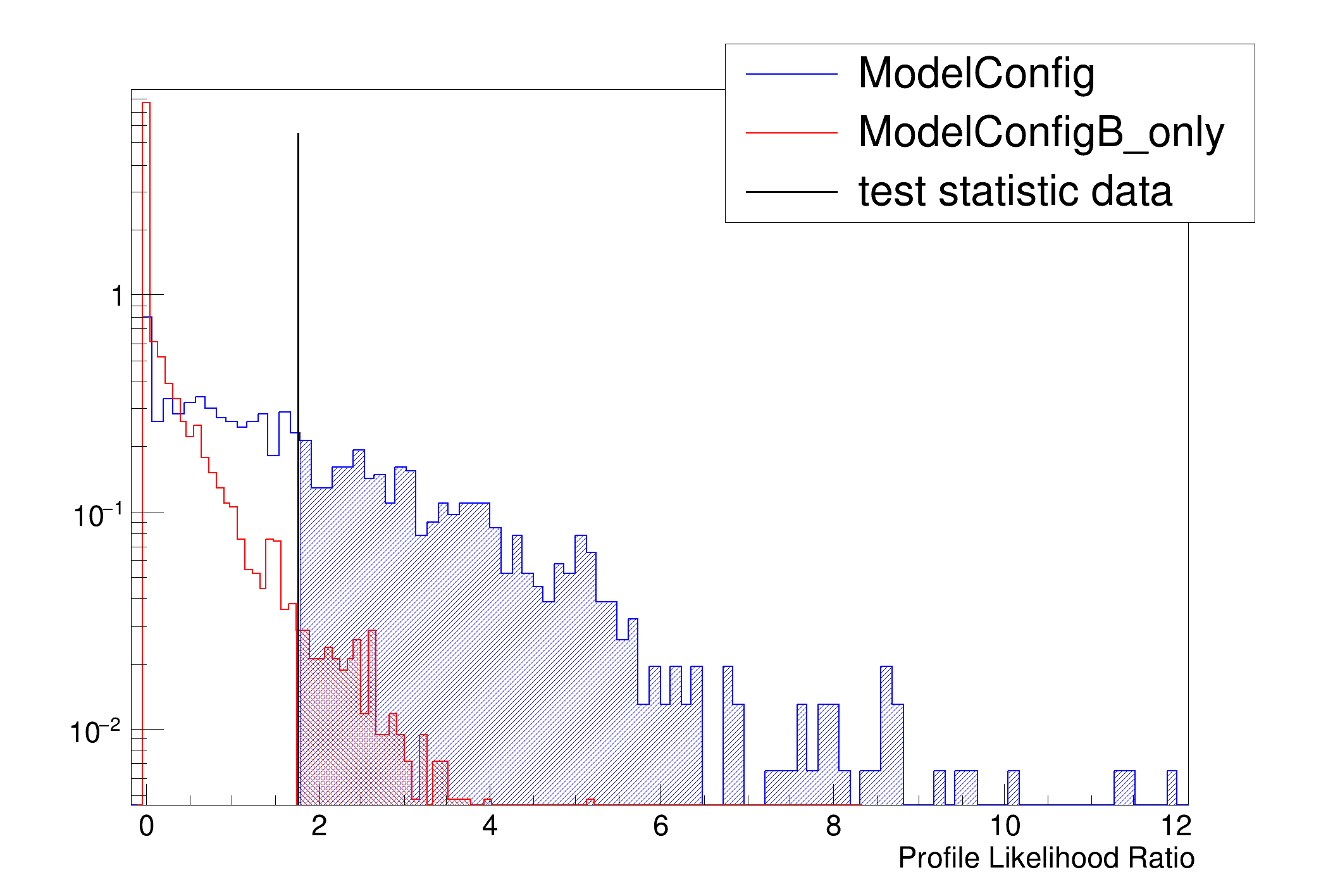
[#0] PROGRESS:Generation -- Test Statistic on data: 1.77404
[#1] INFO:InputArguments -- Profiling conditional MLEs for Null.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Null.
[#0] PROGRESS:Generation -- generated toys: 500 / 5000
[#0] PROGRESS:Generation -- generated toys: 1000 / 5000
[#0] PROGRESS:Generation -- generated toys: 1500 / 5000
[#0] PROGRESS:Generation -- generated toys: 2000 / 5000
[#0] PROGRESS:Generation -- generated toys: 2500 / 5000
[#0] PROGRESS:Generation -- generated toys: 3000 / 5000
[#0] PROGRESS:Generation -- generated toys: 3500 / 5000
[#0] PROGRESS:Generation -- generated toys: 4000 / 5000
[#0] PROGRESS:Generation -- generated toys: 4500 / 5000
[#1] INFO:InputArguments -- Profiling conditional MLEs for Alt.
[#1] INFO:InputArguments -- Using a ToyMCSampler. Now configuring for Alt.
[#0] PROGRESS:Generation -- generated toys: 500 / 1250
[#0] PROGRESS:Generation -- generated toys: 1000 / 1250
Results HypoTestCalculator_result:
- Null p-value = 0.0304 +/- 0.002428
- Significance = 1.87495 +/- 0.0352942 sigma
- Number of Alt toys: 1250
- Number of Null toys: 5000
- Test statistic evaluated on data: 1.77404
- CL_b: 0.0304 +/- 0.002428
- CL_s+b: 0.4328 +/- 0.0140138
- CL_s: 14.2368 +/- 1.22696
Expected p -value and significance at -2 sigma = 0.839 significance -0.990356 sigma
Expected p -value and significance at -1 sigma = 0.2238 significance 0.759422 sigma
Expected p -value and significance at 0 sigma = 0.0434 significance 1.71252 sigma
Expected p -value and significance at 1 sigma = 0.0028 significance 2.77033 sigma
Expected p -value and significance at 2 sigma = 0.0002 significance 3.54008 sigma
#include <cassert>
struct HypoTestOptions {
bool noSystematics = false;
double nToysRatio = 4;
double poiValue = -1;
int printLevel=0;
bool generateBinned = false;
bool useProof = false;
bool enableDetailedOutput = false;
};
HypoTestOptions optHT;
void StandardHypoTestDemo(const char* infile = "",
const char* workspaceName = "combined",
const char* modelSBName = "ModelConfig",
const char* modelBName = "",
const char* dataName = "obsData",
int calcType = 0,
int testStatType = 3,
int ntoys = 5000,
bool useNC = false,
const char * nuisPriorName = 0)
{
bool noSystematics = optHT.noSystematics;
double nToysRatio = optHT.nToysRatio;
double poiValue = optHT.poiValue;
int printLevel = optHT.printLevel;
bool generateBinned = optHT.generateBinned;
bool useProof = optHT.useProof;
bool enableDetOutput = optHT.enableDetailedOutput;
SimpleLikelihoodRatioTestStat::SetAlwaysReuseNLL(true);
ProfileLikelihoodTestStat::SetAlwaysReuseNLL(true);
RatioOfProfiledLikelihoodsTestStat::SetAlwaysReuseNLL(true);
const char* filename = "";
if (!strcmp(infile,"")) {
filename = "results/example_combined_GaussExample_model.root";
if (!fileExist) {
#ifdef _WIN32
cout << "HistFactory file cannot be generated on Windows - exit" << endl;
return;
#endif
cout <<"will run standard hist2workspace example"<<endl;
gROOT->ProcessLine(
".! prepareHistFactory .");
gROOT->ProcessLine(
".! hist2workspace config/example.xml");
cout <<"\n\n---------------------"<<endl;
cout <<"Done creating example input"<<endl;
cout <<"---------------------\n\n"<<endl;
}
}
else
filename = infile;
cout <<"StandardRooStatsDemoMacro: Input file " << filename << " is not found" << endl;
return;
}
if(!w){
cout <<"workspace not found" << endl;
return;
}
ModelConfig* sbModel = (ModelConfig*) w->
obj(modelSBName);
cout << "data or ModelConfig was not found" <<endl;
return;
}
ModelConfig* bModel = (ModelConfig*) w->
obj(modelBName);
if (noSystematics) {
const RooArgSet * nuisPar = sbModel->GetNuisanceParameters();
if (nuisPar && nuisPar->
getSize() > 0) {
std::cout << "StandardHypoTestInvDemo" << " - Switch off all systematics by setting them constant to their initial values" << std::endl;
}
if (bModel) {
const RooArgSet * bnuisPar = bModel->GetNuisanceParameters();
if (bnuisPar)
}
}
if (!bModel ) {
Info(
"StandardHypoTestInvDemo",
"The background model %s does not exist",modelBName);
Info(
"StandardHypoTestInvDemo",
"Copy it from ModelConfig %s and set POI to zero",modelSBName);
bModel = (ModelConfig*) sbModel->Clone();
if (!var) return;
double oldval = var->
getVal();
}
if (!sbModel->GetSnapshot() || poiValue > 0) {
Info(
"StandardHypoTestDemo",
"Model %s has no snapshot - make one using model poi",modelSBName);
if (!var) return;
double oldval = var->
getVal();
if (poiValue > 0) var->
setVal(poiValue);
if (poiValue > 0) var->
setVal(oldval);
}
SimpleLikelihoodRatioTestStat * slrts = new SimpleLikelihoodRatioTestStat(*bModel->GetPdf(), *sbModel->GetPdf());
RooArgSet nullParams(*bModel->GetSnapshot());
if (bModel->GetNuisanceParameters()) nullParams.add(*bModel->GetNuisanceParameters());
slrts->SetNullParameters(nullParams);
RooArgSet altParams(*sbModel->GetSnapshot());
if (sbModel->GetNuisanceParameters()) altParams.add(*sbModel->GetNuisanceParameters());
slrts->SetAltParameters(altParams);
ProfileLikelihoodTestStat * profll = new ProfileLikelihoodTestStat(*bModel->GetPdf());
RatioOfProfiledLikelihoodsTestStat *
ropl = new RatioOfProfiledLikelihoodsTestStat(*bModel->GetPdf(), *sbModel->GetPdf(), sbModel->GetSnapshot());
ropl->SetSubtractMLE(false);
if (testStatType == 3) profll->SetOneSidedDiscovery(1);
profll->SetPrintLevel(printLevel);
if (enableDetOutput) {
slrts->EnableDetailedOutput();
profll->EnableDetailedOutput();
ropl->EnableDetailedOutput();
}
AsymptoticCalculator::SetPrintLevel(printLevel);
HypoTestCalculatorGeneric * hypoCalc = 0;
if (calcType == 0) hypoCalc =
new FrequentistCalculator(*
data, *sbModel, *bModel);
else if (calcType == 1) hypoCalc=
new HybridCalculator(*
data, *sbModel, *bModel);
else if (calcType == 2) hypoCalc=
new AsymptoticCalculator(*
data, *sbModel, *bModel);
if (calcType == 0) {
((FrequentistCalculator*)hypoCalc)->SetToys(ntoys, ntoys/nToysRatio);
if (enableDetOutput) ((FrequentistCalculator*) hypoCalc)->StoreFitInfo(true);
}
if (calcType == 1) {
((HybridCalculator*)hypoCalc)->SetToys(ntoys, ntoys/nToysRatio);
}
if (calcType == 2 ) {
if (testStatType == 3) ((AsymptoticCalculator*) hypoCalc)->SetOneSidedDiscovery(true);
if (testStatType != 2 && testStatType != 3)
Warning(
"StandardHypoTestDemo",
"Only the PL test statistic can be used with AsymptoticCalculator - use by default a two-sided PL");
}
if (calcType == 1 && (bModel->GetNuisanceParameters() || sbModel->GetNuisanceParameters() )) {
if (nuisPriorName) nuisPdf = w->
pdf(nuisPriorName);
if (!nuisPdf) {
Info(
"StandardHypoTestDemo",
"No nuisance pdf given for the HybridCalculator - try to deduce pdf from the model");
if (bModel->GetPdf() && bModel->GetObservables() )
else
}
if (!nuisPdf ) {
if (bModel->GetPriorPdf()) {
nuisPdf = bModel->GetPriorPdf();
Info(
"StandardHypoTestDemo",
"No nuisance pdf given - try to use %s that is defined as a prior pdf in the B model",nuisPdf->
GetName());
}
else {
Error(
"StandardHypoTestDemo",
"Cannot run Hybrid calculator because no prior on the nuisance parameter is specified or can be derived");
return;
}
}
assert(nuisPdf);
Info(
"StandardHypoTestDemo",
"Using as nuisance Pdf ... " );
const RooArgSet * nuisParams = (bModel->GetNuisanceParameters() ) ? bModel->GetNuisanceParameters() : sbModel->GetNuisanceParameters();
Warning(
"StandardHypoTestDemo",
"Prior nuisance does not depend on nuisance parameters. They will be smeared in their full range");
}
delete np;
((HybridCalculator*)hypoCalc)->ForcePriorNuisanceAlt(*nuisPdf);
((HybridCalculator*)hypoCalc)->ForcePriorNuisanceNull(*nuisPdf);
}
ToyMCSampler * sampler = (ToyMCSampler *)hypoCalc->GetTestStatSampler();
if (sampler && (calcType == 0 || calcType == 1) ) {
if (sbModel->GetPdf()->canBeExtended() ) {
if (useNC)
Warning(
"StandardHypoTestDemo",
"Pdf is extended: but number counting flag is set: ignore it ");
}
else {
if (!useNC) {
int nEvents =
data->numEntries();
Info(
"StandardHypoTestDemo",
"Pdf is not extended: number of events to generate taken from observed data set is %d",nEvents);
sampler->SetNEventsPerToy(nEvents);
}
else {
Info(
"StandardHypoTestDemo",
"using a number counting pdf");
sampler->SetNEventsPerToy(1);
}
}
if (
data->isWeighted() && !generateBinned) {
Info(
"StandardHypoTestDemo",
"Data set is weighted, nentries = %d and sum of weights = %8.1f but toy generation is unbinned - it would be faster to set generateBinned to true\n",
data->numEntries(),
data->sumEntries());
}
if (generateBinned) sampler->SetGenerateBinned(generateBinned);
if (useProof) {
sampler->SetProofConfig(&
pc);
}
if (testStatType == 0) sampler->SetTestStatistic(slrts);
if (testStatType == 1) sampler->SetTestStatistic(ropl);
if (testStatType == 2 || testStatType == 3) sampler->SetTestStatistic(profll);
}
HypoTestResult * htr = hypoCalc->GetHypoTest();
htr->SetPValueIsRightTail(true);
htr->SetBackgroundAsAlt(false);
htr->Print();
delete sampler;
delete slrts;
delete ropl;
delete profll;
if (calcType != 2) {
HypoTestPlot * plot = new HypoTestPlot(*htr,100);
plot->SetLogYaxis(true);
plot->Draw();
}
else {
std::cout << "Asymptotic results " << std::endl;
}
if (calcType != 2) {
SamplingDistribution * altDist = htr->GetAltDistribution();
HypoTestResult htExp("Expected Result");
htExp.Append(htr);
double p[5];
for (int i = 0; i < 5; ++i) {
double sig = -2 + i;
}
std::vector<double> values = altDist->GetSamplingDistribution();
for (int i = 0; i < 5; ++i) {
htExp.SetTestStatisticData(
q[i] );
double sig = -2 + i;
std::cout << " Expected p -value and significance at " << sig << " sigma = "
<< htExp.NullPValue() << " significance " << htExp.Significance() << " sigma " << std::endl;
}
}
else {
for (int i = 0; i < 5; ++i) {
double sig = -2 + i;
double pval = AsymptoticCalculator::GetExpectedPValues( htr->NullPValue(), htr->AlternatePValue(), -sig, false);
std::cout << " Expected p -value and significance at " << sig << " sigma = "
}
}
bool writeResult = (calcType != 2);
if (enableDetOutput) {
writeResult=true;
Info(
"StandardHypoTestDemo",
"Detailed output will be written in output result file");
}
if (htr != NULL && writeResult) {
const char * calcTypeName = (calcType == 0) ? "Freq" : (calcType == 1) ? "Hybr" : "Asym";
TFile * fileOut =
new TFile(resultFileName,
"RECREATE");
htr->Write();
Info(
"StandardHypoTestDemo",
"HypoTestResult has been written in the file %s",resultFileName.
Data());
}
}
void Info(const char *location, const char *msgfmt,...)
void Error(const char *location, const char *msgfmt,...)
void Warning(const char *location, const char *msgfmt,...)
R__EXTERN TSystem * gSystem
RooArgSet * getObservables(const RooArgSet &set, Bool_t valueOnly=kTRUE) const
virtual void Print(Option_t *options=0) const
Print TNamed name and title.
RooAbsData is the common abstract base class for binned and unbinned datasets.
RooAbsPdf is the abstract interface for all probability density functions The class provides hybrid a...
Double_t getVal(const RooArgSet *set=0) const
Evaluate object. Returns either cached value or triggers a recalculation.
RooArgSet is a container object that can hold multiple RooAbsArg objects.
RooRealVar represents a fundamental (non-derived) real valued object.
virtual void setVal(Double_t value)
Set value of variable to 'value'.
The RooWorkspace is a persistable container for RooFit projects.
RooAbsData * data(const char *name) const
Retrieve dataset (binned or unbinned) with given name. A null pointer is returned if not found.
void Print(Option_t *opts=0) const
Print contents of the workspace.
TObject * obj(const char *name) const
Return any type of object (RooAbsArg, RooAbsData or generic object) with given name)
RooAbsPdf * pdf(const char *name) const
Retrieve p.d.f (RooAbsPdf) with given name. A null pointer is returned if not found.
A ROOT file is a suite of consecutive data records (TKey instances) with a well defined format.
virtual void Close(Option_t *option="")
Close a file.
static TFile * Open(const char *name, Option_t *option="", const char *ftitle="", Int_t compress=ROOT::RCompressionSetting::EDefaults::kUseGeneralPurpose, Int_t netopt=0)
Create / open a file.
virtual const char * GetName() const
Returns name of object.
const char * Data() const
static TString Format(const char *fmt,...)
Static method which formats a string using a printf style format descriptor and return a TString.
virtual Bool_t AccessPathName(const char *path, EAccessMode mode=kFileExists)
Returns FALSE if one can access a file using the specified access mode.
double normal_cdf(double x, double sigma=1, double x0=0)
Cumulative distribution function of the normal (Gaussian) distribution (lower tail).
double normal_quantile_c(double z, double sigma)
Inverse ( ) of the cumulative distribution function of the upper tail of the normal (Gaussian) distri...
@(#)root/roostats:$Id$ Author: George Lewis, Kyle Cranmer
bool SetAllConstant(const RooAbsCollection &coll, bool constant=true)
RooAbsPdf * MakeNuisancePdf(RooAbsPdf &pdf, const RooArgSet &observables, const char *name)
static constexpr double pc
void Quantiles(Int_t n, Int_t nprob, Double_t *x, Double_t *quantiles, Double_t *prob, Bool_t isSorted=kTRUE, Int_t *index=0, Int_t type=7)
Computes sample quantiles, corresponding to the given probabilities.



 Standard tutorial macro for hypothesis test (for computing the discovery significance) using all RooStats hypotheiss tests calculators and test statistics.
Standard tutorial macro for hypothesis test (for computing the discovery significance) using all RooStats hypotheiss tests calculators and test statistics.