using namespace std;
TH1 *hfit = (
TH1*)gFitter->GetObjectFit();
TF1 *f1 = (TF1*)gFitter->GetUserFunc();
f1->InitArgs(&x,u);
npar = f1->GetNpar();
f = 0;
for (
Int_t i=0;i<nPoints;i++) {
else npfit++;
}
for (
Int_t i=0;i<nPoints;i++) {
for (
Int_t j=0;j<nPoints;j++) {
f += df[i]*df[j]*gHistInvEMatrix->GetBinContent(i+1,j+1);
}
}
delete[] df;
f1->SetNumberFitPoints(npfit);
}
return par[0]/(dm*dm+par[2]*par[2]);
}
{
if(rnd->Rndm()>bgr) {
do {
do {
t=rnd->Rndm();
} while(t>=1.0);
} while(t<=0.0);
} else {
do {
do {
t=rnd->Rndm();
} while(t>=1.0);
} while(t>=0.0);
}
}
if(rnd->Rndm()>frac) {
return rnd->Gaus(mTrue+smallBias,smallSigma);
} else {
return rnd->Gaus(mTrue+wideBias,wideSigma);
}
}
int testUnfold1()
{
TH1D *histMgenMC=
new TH1D(
"MgenMC",
";mass(gen)",nGen,xminGen,xmaxGen);
TH1D *histMdetMC=
new TH1D(
"MdetMC",
";mass(det)",nDet,xminDet,xmaxDet);
TH2D *histMdetGenMC=
new TH2D(
"MdetgenMC",
";mass(det);mass(gen)",
nDet,xminDet,xmaxDet,nGen,xminGen,xmaxGen);
Int_t neventMC=rnd->Poisson(luminosityMC*crossSection);
for(
Int_t i=0;i<neventMC;i++) {
4.0,
0.2);
histMgenMC->
Fill(mGen,luminosityData/luminosityMC);
histMdetMC->
Fill(mDet,luminosityData/luminosityMC);
histMdetGenMC->
Fill(mDet,mGen,luminosityData/luminosityMC);
}
TH2D *histMdetGenSysMC=
new TH2D(
"MdetgenSysMC",
";mass(det);mass(gen)",
nDet,xminDet,xmaxDet,nGen,xminGen,xmaxGen);
neventMC=rnd->Poisson(luminosityMC*crossSection);
for(
Int_t i=0;i<neventMC;i++) {
(0.5,
3.6,
0.15);
histMdetGenSysMC->
Fill(mDet,mGen,luminosityData/luminosityMC);
}
TH1D *histMgenData=
new TH1D(
"MgenData",
";mass(gen)",nGen,xminGen,xmaxGen);
TH1D *histMdetData=
new TH1D(
"MdetData",
";mass(det)",nDet,xminDet,xmaxDet);
Int_t neventData=rnd->Poisson(luminosityData*crossSection);
for(
Int_t i=0;i<neventData;i++) {
3.8,
0.15);
histMgenData->
Fill(mGen);
histMdetData->
Fill(mDet);
}
TH1D *histDensityGenData=
new TH1D(
"DensityGenData",
";mass(gen)",
nGen,xminGen,xmaxGen);
TH1D *histDensityGenMC=
new TH1D(
"DensityGenMC",
";mass(gen)",
nGen,xminGen,xmaxGen);
for(
Int_t i=1;i<=nGen;i++) {
}
if(unfold.SetInput(histMdetData)>=10000) {
std::cout<<"Unfolding result may be wrong\n";
}
TGraph *lCurve;
#ifdef VERBOSE_LCURVE_SCAN
#endif
iBest=unfold.ScanLcurve(nScan,tauMin,tauMax,&lCurve,&logTauX,&logTauY);
#ifdef VERBOSE_LCURVE_SCAN
#endif
TH2D *histMdetGenSys1=
new TH2D(
"Mdetgensys1",
";mass(det);mass(gen)",
nDet,xminDet,xmaxDet,nGen,xminGen,xmaxGen);
for(
Int_t i=0;i<=nDet+1;i++) {
for(
Int_t j=0;j<=nGen+1;j++) {
}
}
}
std::cout<<"tau="<<unfold.GetTau()<<"\n";
std::cout<<"chi**2="<<unfold.GetChi2A()<<"+"<<unfold.GetChi2L()
<<" / "<<unfold.GetNdf()<<"\n";
std::cout<<"chi**2(sys)="<<unfold.GetChi2Sys()<<"\n";
TGraph *bestLcurve=new TGraph(1,x,y);
TGraph *bestLogTauLogChi2=new TGraph(1,t,x);
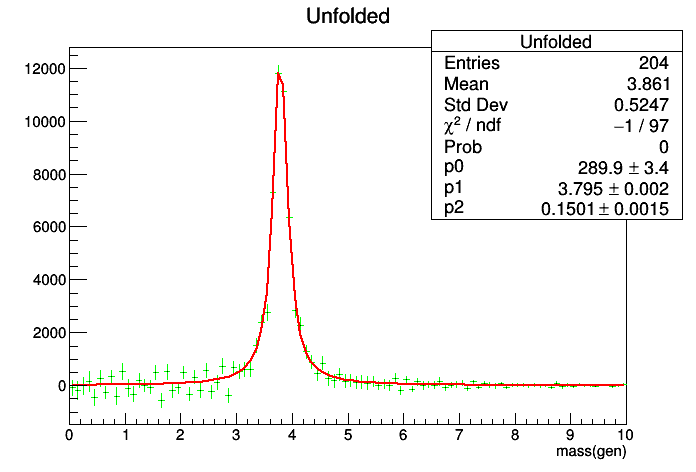
TH1 *histMunfold=unfold.GetOutput(
"Unfolded");
TH1 *histMdetFold=unfold.GetFoldedOutput(
"FoldedBack");
TH2 *histEmatTotal=unfold.GetEmatrixTotal(
"EmatTotal");
new TH1D(
"TotalError",
";mass(gen)",nGen,xminGen,xmaxGen);
for(
Int_t bin=1;bin<=nGen;bin++) {
}
TH1 *histRhoi=unfold.GetRhoItotal(
"rho_I",
0,
0,
"*[UO]",
&gHistInvEMatrix
);
gFitter->SetFCN(chisquare_corr);
TF1 *bw=new TF1("bw",bw_func,xminGen,xmaxGen,3);
bw->SetParameter(0,1000.);
bw->SetParameter(1,3.8);
bw->SetParameter(2,0.2);
histMunfold->
Fit(bw,
"UE");
output.Divide(3,2);
output.cd(1);
histMdetGenMC->
Draw(
"BOX");
output.cd(2);
histTotalError->
Draw(
"E");
histMunfold->
Draw(
"SAME E1");
histDensityGenData->
Draw(
"SAME");
histDensityGenMC->
Draw(
"SAME HIST");
output.cd(3);
histMdetMC->
Draw(
"SAME HIST");
TH1 *histInput=unfold.GetInput(
"Minput",
";mass(det)");
output.cd(4);
output.cd(5);
bestLogTauLogChi2->SetMarkerColor(
kRed);
bestLogTauLogChi2->Draw("*");
output.cd(6);
lCurve->Draw("AL");
bestLcurve->SetMarkerColor(
kRed);
bestLcurve->Draw("*");
output.SaveAs("testUnfold1.ps");
return 0;
}