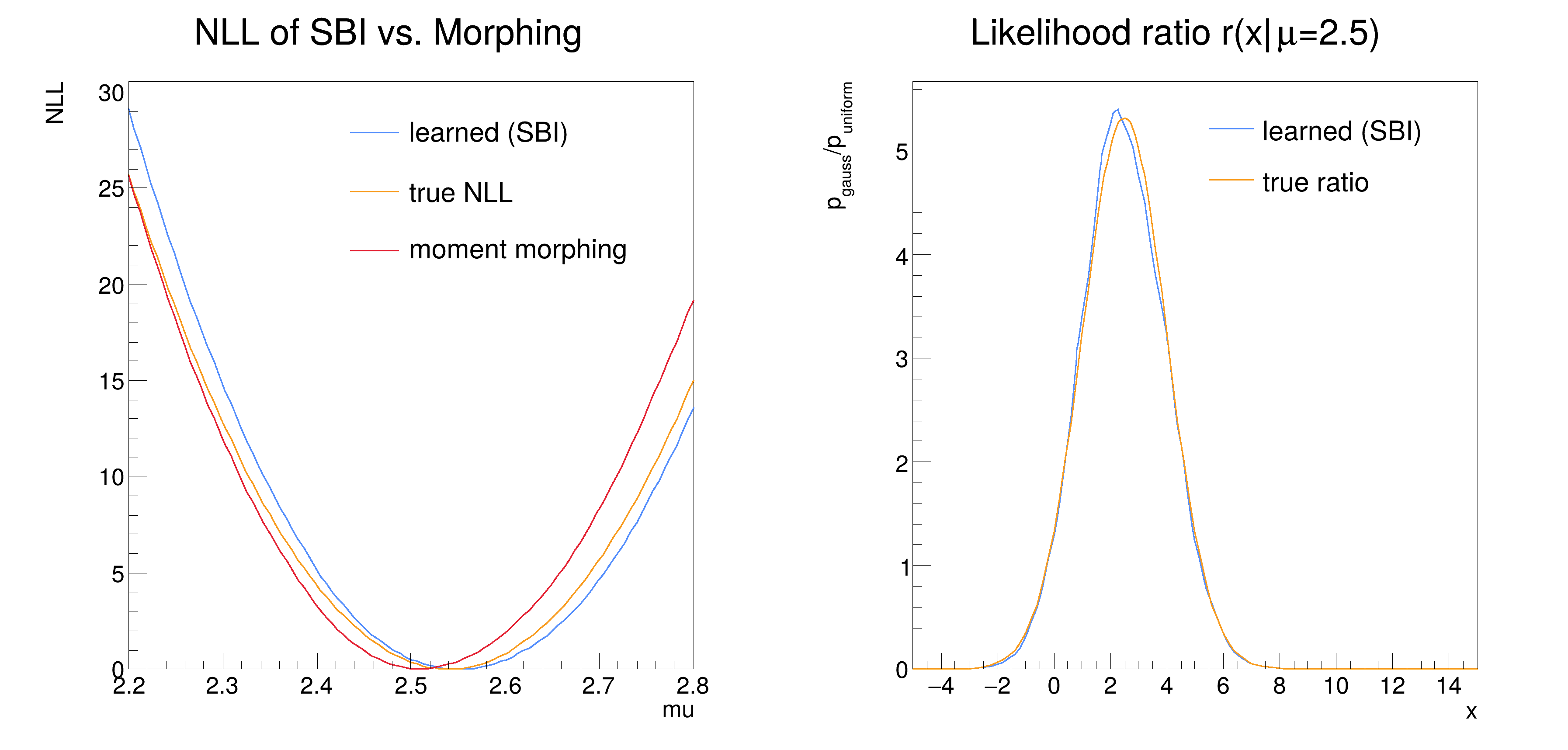
We compare the approach of using the likelihood ratio trick to morphing.
A short recap: The idea of SBI is to fit a surrogate model to the data, in order to really learn the likelihood function instead of calculating it. Therefore, a classifier is trained to discriminate between samples from a target distribution (here the Gaussian) $$x\sim p(x|\theta)$$ and a reference distribution (here the Uniform) $$x\sim p_{ref}(x|\theta)$$.
The output of the classifier $$\hat{s}(\theta)$$ is a monotonic function of the likelihood ration and can be turned into an estimate of the likelihood ratio via $$\hat{r}(\theta)=\frac{1-\hat{s}(\theta)}{\hat{s}(\theta)}.$$ This is called the likelihood ratio trick.
In the end we compare the negative logarithmic likelihoods of the learned, morphed and analytical likelihood with minuit and as a plot.
import numpy as np
import ROOT
n_samples = 10000
n_grid = 5
hist = workspace[f"g{i}"].generateBinned([x_var], n_samples)
def __init__(self, workspace):
self.classifier =
MLPClassifier(hidden_layer_sizes=(20, 20), max_iter=1000, random_state=42)
self.data_model = None
self.data_ref = None
self.X_train = None
self.y_train = None
self.workspace = workspace
ws = self.workspace
data_test_model = []
samples_gaussian = ws[model].generate([ws[x], ws[mu]], n_samples).
to_numpy()
self._training_mus = samples_gaussian[mu]
ws = self.workspace
samples_uniform = ws[model].generate(ws[x], n_samples)
)
thetas_model = self._training_mus.
reshape(-1, 1)
thetas_reference = self._training_mus.
reshape(-1, 1)
self.classifier.fit(self.X_train, self.y_train)
n_samples_train = n_samples * 5
ws.factory(f
"Gaussian::gauss(x[-5,15], mu[0,4], {sigma})")
ws["mu"].setVal(mu_observed)
obs_data = ws["gauss"].generate(ws["x"], 1000)
return ws
mu_observed = 2.5
sigma = 1.5
workspace =
build_ws(mu_observed, sigma)
x_var = workspace["x"]
mu_var = workspace["mu"]
gauss = workspace["gauss"]
uniform = workspace["uniform"]
obs_data = workspace["obs_data"]
sbi_model = model
X[:, 0] = x
X[:, 1] = mu
return prob / (1 - prob)
nll_morph = workspace["morph"].createNLL(obs_data)
frame1 =
mu_var.frame(Title=
"NLL of SBI vs. Morphing;mu;NLL", Range=(2.2, 2.8))
frame2 =
x_var.frame(Title=
"Likelihood ratio r(x|#mu=2.5);x;p_{gauss}/p_{uniform}")
single_canvas = True
c =
ROOT.TCanvas(
"",
"", 1200
if single_canvas
else 600, 600)
if single_canvas:
if single_canvas:
else:
if not single_canvas:
for nll in [nll_gauss, nllr_learned, nll_morph]:
del nll_morph
del nllr_learned
del nll_gauss
del workspace
ROOT::Detail::TRangeCast< T, true > TRangeDynCast
TRangeDynCast is an adapter class that allows the typed iteration through a TCollection.
Option_t Option_t TPoint TPoint const char GetTextMagnitude GetFillStyle GetLineColor GetLineWidth GetMarkerStyle GetTextAlign GetTextColor GetTextSize void char Point_t Rectangle_t WindowAttributes_t Float_t Float_t Float_t Int_t Int_t UInt_t UInt_t Rectangle_t Int_t Int_t Window_t TString Int_t GCValues_t GetPrimarySelectionOwner GetDisplay GetScreen GetColormap GetNativeEvent const char const char dpyName wid window const char font_name cursor keysym reg const char only_if_exist regb h Point_t winding char text const char depth char const char Int_t count const char ColorStruct_t color const char Pixmap_t Pixmap_t PictureAttributes_t attr const char char ret_data h unsigned char height h Atom_t Int_t ULong_t ULong_t unsigned char prop_list Atom_t Atom_t Atom_t Time_t UChar_t len
RooWorkspace() contents
variables
---------
(mu,x)
p.d.f.s
-------
RooGaussian::gauss[ x=x mean=mu sigma=1.5 ] = 0.249352
RooUniform::uniform[ x=(x) ] = 1
[#0] ERROR:Eval -- RooAbsReal::logEvalError(morph_func_offset_0_0) evaluation error,
origin : RooFormulaVar::morph_func_offset_0_0[ actualVars=(histpdf0_MOMENT_1_x,morph_func_pos_0,morph_func_slope_0_0) formula="x[0]-(x[1]*x[2])" ]
message : function value is NAN
server values: actualVars=(histpdf0_MOMENT_1_x = 0.0504296,morph_func_pos_0 = 0,morph_func_slope_0_0 = inf)
[#0] ERROR:Eval -- RooAbsReal::logEvalError(morph_func_offset_1_0) evaluation error,
origin : RooFormulaVar::morph_func_offset_1_0[ actualVars=(histpdf1_MOMENT_1_x,morph_func_pos_0,morph_func_slope_1_0) formula="x[0]-(x[1]*x[2])" ]
message : function value is NAN
server values: actualVars=(histpdf1_MOMENT_1_x = 0.997298,morph_func_pos_0 = 0,morph_func_slope_1_0 = inf)
[#0] ERROR:Eval -- RooAbsReal::logEvalError(morph_func_offset_2_0) evaluation error,
origin : RooFormulaVar::morph_func_offset_2_0[ actualVars=(histpdf2_MOMENT_1_x,morph_func_pos_0,morph_func_slope_2_0) formula="x[0]-(x[1]*x[2])" ]
message : function value is NAN
server values: actualVars=(histpdf2_MOMENT_1_x = 2.01481,morph_func_pos_0 = 0,morph_func_slope_2_0 = inf)
[#0] ERROR:Eval -- RooAbsReal::logEvalError(morph_func_offset_3_0) evaluation error,
origin : RooFormulaVar::morph_func_offset_3_0[ actualVars=(histpdf3_MOMENT_1_x,morph_func_pos_0,morph_func_slope_3_0) formula="x[0]-(x[1]*x[2])" ]
message : function value is NAN
server values: actualVars=(histpdf3_MOMENT_1_x = 3.04583,morph_func_pos_0 = 0,morph_func_slope_3_0 = inf)
[#0] ERROR:Eval -- RooAbsReal::logEvalError(morph_func_offset_4_0) evaluation error,
origin : RooFormulaVar::morph_func_offset_4_0[ actualVars=(histpdf4_MOMENT_1_x,morph_func_pos_0,morph_func_slope_4_0) formula="x[0]-(x[1]*x[2])" ]
message : function value is NAN
server values: actualVars=(histpdf4_MOMENT_1_x = 4.00616,morph_func_pos_0 = 0,morph_func_slope_4_0 = inf)
RooFitResult: minimized FCN value: 1862.97, estimated distance to minimum: 2.32702e-05
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
mu 2.5399e+00 +/- 4.74e-02
RooFitResult: minimized FCN value: -1126.14, estimated distance to minimum: 4.23342e-05
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
mu 2.5511e+00 +/- 7.15e-02
RooFitResult: minimized FCN value: -7350.75, estimated distance to minimum: 0.000125595
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
mu 2.5103e+00 +/- 4.08e-02