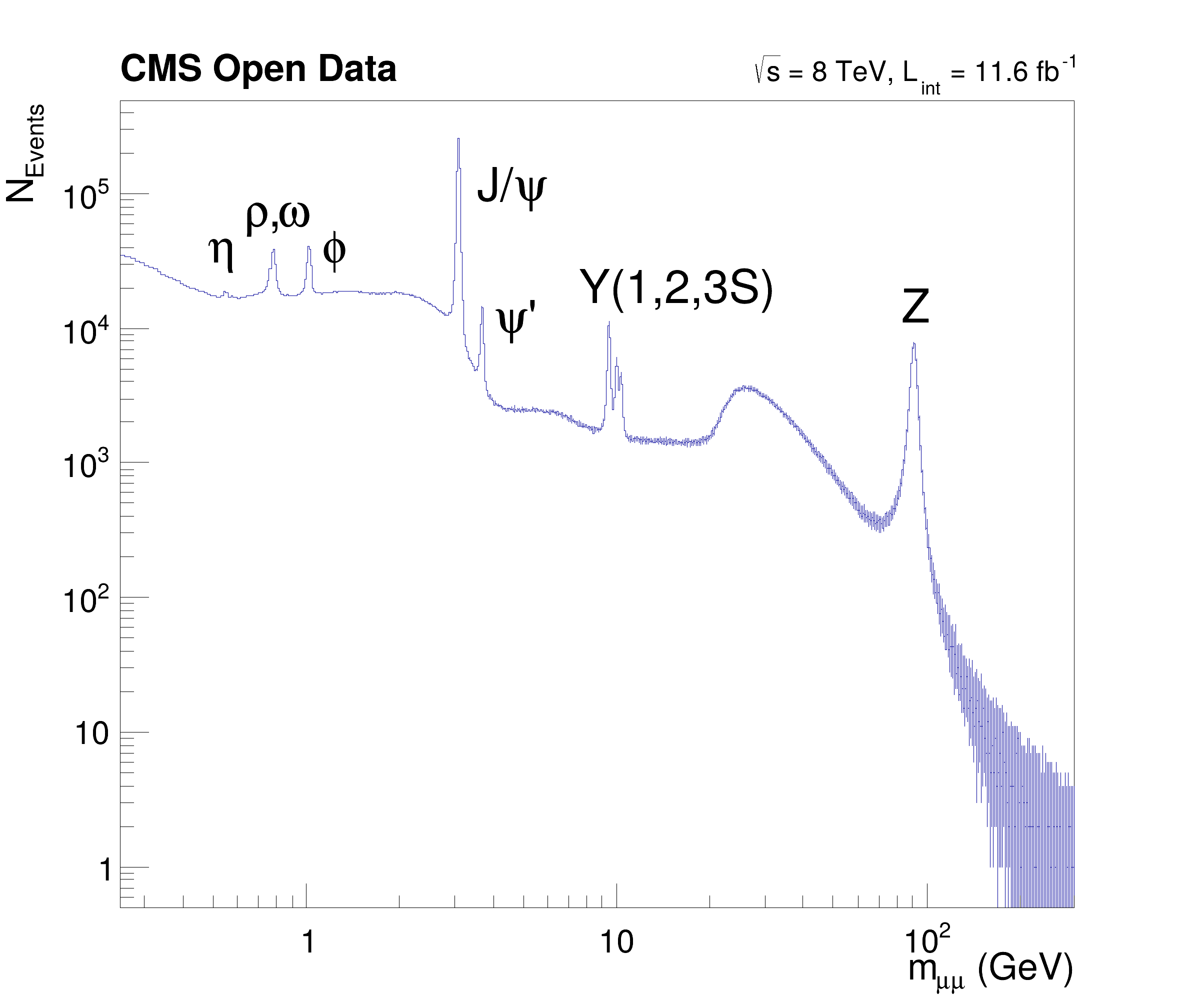
Show how NanoAOD files can be processed with RDataFrame.
This tutorial illustrates how NanoAOD files can be processed with ROOT dataframes. The NanoAOD-like input files are filled with 66 mio. events from CMS OpenData containing muon candidates part of 2012 dataset (DOI: 10.7483/OPENDATA.CMS.YLIC.86ZZ and DOI: 10.7483/OPENDATA.CMS.M5AD.Y3V3). The macro matches muon pairs and produces an histogram of the dimuon mass spectrum showing resonances up to the Z mass. Note that the bump at 30 GeV is not a resonance but a trigger effect.
More details about the dataset can be found on the CERN Open Data portal.
import ROOT
df =
ROOT.RDataFrame(
"Events",
"root://eospublic.cern.ch//eos/opendata/cms/derived-data/AOD2NanoAODOutreachTool/Run2012BC_DoubleMuParked_Muons.root")
df_2mu =
df.Filter(
"nMuon == 2",
"Events with exactly two muons")
df_os =
df_2mu.Filter(
"Muon_charge[0] != Muon_charge[1]",
"Muons with opposite charge")
df_mass =
df_os.Define(
"Dimuon_mass",
"InvariantMass(Muon_pt, Muon_eta, Muon_phi, Muon_mass)")
h =
df_mass.Histo1D((
"Dimuon_mass",
"Dimuon mass;m_{#mu#mu} (GeV);N_{Events}", 30000, 0.25, 300),
"Dimuon_mass")
ROOT::Detail::TRangeCast< T, true > TRangeDynCast
TRangeDynCast is an adapter class that allows the typed iteration through a TCollection.
ROOT's RDataFrame offers a modern, high-level interface for analysis of data stored in TTree ,...
Events with exactly two muons: pass=31104343 all=61540413 -- eff=50.54 % cumulative eff=50.54 %
Muons with opposite charge: pass=24067843 all=31104343 -- eff=77.38 % cumulative eff=39.11 %
- Date
- April 2019
- Author
- Stefan Wunsch (KIT, CERN)
Definition in file df102_NanoAODDimuonAnalysis.py.