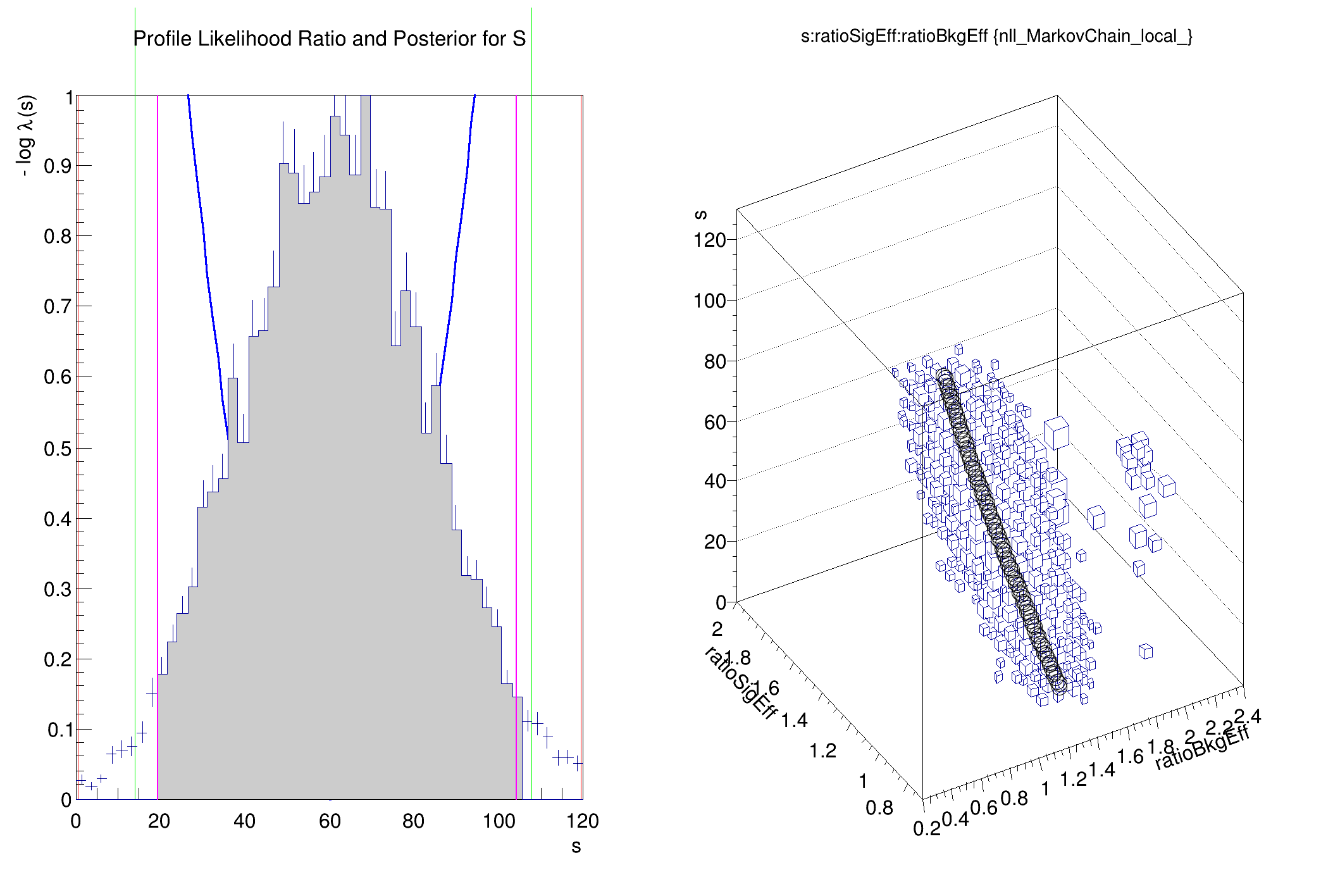
The usage of a Confidence Interval Calculator to set a limit on the signal is illustrated
RooWorkspace() contents
variables
---------
(b,gSigBkg,gSigEff,obs,ratioBkgEff,ratioSigEff,s)
p.d.f.s
-------
RooGaussian::bkgConstraint[ x=gSigBkg mean=ratioBkgEff sigma=0.2 ] = 1
RooPoisson::countingModel[ x=obs mean=countingModel_2 ] = 0.0325554
RooProdPdf::modelWithConstraints[ countingModel * sigConstraint * bkgConstraint ] = 0.0325554
RooGaussian::sigConstraint[ x=gSigEff mean=ratioSigEff sigma=0.05 ] = 1
functions
--------
RooAddition::countingModel_2[ countingModel_2_[s_x_ratioSigEff] + countingModel_2_[b_x_ratioBkgEff] ] = 150
RooProduct::countingModel_2_[b_x_ratioBkgEff][ b * ratioBkgEff ] = 100
RooProduct::countingModel_2_[s_x_ratioSigEff][ s * ratioSigEff ] = 50
[#1] INFO:Minimization -- Including the following constraint terms in minimization: (sigConstraint,bkgConstraint)
[#1] INFO:Minimization -- The global observables are not defined , normalize constraints with respect to the parameters (ratioSigEff,ratioBkgEff)
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_modelWithConstraints_exampleData_with_constr) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (countingModel)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
[#1] INFO:ObjectHandling -- RooWorkspace::import() importing dataset exampleData
[#1] INFO:InputArguments -- The deprecated RooFit::CloneData(1) option passed to createNLL() is ignored.
[#1] INFO:Minimization -- Including the following constraint terms in minimization: (sigConstraint,bkgConstraint)
[#1] INFO:Minimization -- The following global observables have been defined and their values are taken from the model: (gSigEff,gSigBkg)
[#0] PROGRESS:Minimization -- ProfileLikelihoodCalcultor::DoGLobalFit - find MLE
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_modelWithConstraints_exampleData_with_constr) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (countingModel)
[#0] PROGRESS:Minimization -- ProfileLikelihoodCalcultor::DoMinimizeNLL - using Minuit2 / Migrad with strategy 1
[#1] INFO:Minimization --
RooFitResult: minimized FCN value: 0.689753, estimated distance to minimum: 3.0067e-16
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
ratioBkgEff 1.0000e+00 +/- 1.99e-01
ratioSigEff 1.0000e+00 +/- 5.00e-02
s 6.0000e+01 +/- 2.32e+01
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) Creating instance of MINUIT
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_modelWithConstraints_exampleData_with_constr) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) determining minimum likelihood for current configurations w.r.t all observable
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) minimum found at (s=60)
.
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) Creating instance of MINUIT
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_modelWithConstraints_exampleData_with_constr) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) determining minimum likelihood for current configurations w.r.t all observable
[#0] ERROR:InputArguments -- RooArgSet::checkForDup: ERROR argument with name s is already in this set
[#1] INFO:Minimization -- RooProfileLL::evaluate(nll_modelWithConstraints_exampleData_with_constr_Profile[s]) minimum found at (s=60)
..........................................................................................................................................................................................................
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs)
Parameters of Interest: RooArgSet:: = (s)
Nuisance Parameters: RooArgSet:: = (ratioSigEff,ratioBkgEff)
Global Observables: RooArgSet:: = (gSigEff,gSigBkg)
PDF: RooProdPdf::modelWithConstraints[ countingModel * sigConstraint * bkgConstraint ] = 0.00467703
FeldmanCousins: ntoys per point: adaptive
FeldmanCousins: nEvents per toy will not fluctuate, will always be 1
FeldmanCousins: Model has nuisance parameters, will do profile construction
FeldmanCousins: # points to test = 100
NeymanConstruction: Prog: 1/100 total MC = 80 this test stat = 3.21178
s=0.6 ratioSigEff=0.999977 ratioBkgEff=1.43644 [-inf, 4.68588] in interval = 1
NeymanConstruction: Prog: 2/100 total MC = 80 this test stat = 3.08184
s=1.8 ratioSigEff=1.00018 ratioBkgEff=1.42695 [-inf, 3.7189] in interval = 1
NeymanConstruction: Prog: 3/100 total MC = 80 this test stat = 2.95515
s=3 ratioSigEff=1.00055 ratioBkgEff=1.4177 [-inf, 3.91249] in interval = 1
NeymanConstruction: Prog: 4/100 total MC = 80 this test stat = 2.83091
s=4.2 ratioSigEff=1.00106 ratioBkgEff=1.40849 [-inf, 3.65947] in interval = 1
NeymanConstruction: Prog: 5/100 total MC = 80 this test stat = 2.7092
s=5.4 ratioSigEff=1.00133 ratioBkgEff=1.39956 [-inf, 3.10323] in interval = 1
NeymanConstruction: Prog: 6/100 total MC = 80 this test stat = 2.59009
s=6.6 ratioSigEff=1.00159 ratioBkgEff=1.39061 [-inf, 2.97611] in interval = 1
NeymanConstruction: Prog: 7/100 total MC = 240 this test stat = 2.47431
s=7.8 ratioSigEff=1.00103 ratioBkgEff=1.38085 [-inf, 3.25381] in interval = 1
NeymanConstruction: Prog: 8/100 total MC = 80 this test stat = 2.36072
s=9 ratioSigEff=1.00135 ratioBkgEff=1.37176 [-inf, 2.92354] in interval = 1
NeymanConstruction: Prog: 9/100 total MC = 80 this test stat = 2.24982
s=10.2 ratioSigEff=1.00166 ratioBkgEff=1.36273 [-inf, 3.4069] in interval = 1
NeymanConstruction: Prog: 10/100 total MC = 80 this test stat = 2.14164
s=11.4 ratioSigEff=1.00196 ratioBkgEff=1.35375 [-inf, 3.07011] in interval = 1
NeymanConstruction: Prog: 11/100 total MC = 80 this test stat = 2.03623
s=12.6 ratioSigEff=1.00273 ratioBkgEff=1.34407 [-inf, 3.24614] in interval = 1
NeymanConstruction: Prog: 12/100 total MC = 80 this test stat = 1.93333
s=13.8 ratioSigEff=1.00294 ratioBkgEff=1.3353 [-inf, 2.4475] in interval = 1
NeymanConstruction: Prog: 13/100 total MC = 80 this test stat = 1.83331
s=15 ratioSigEff=1.00231 ratioBkgEff=1.32686 [-inf, 2.70017] in interval = 1
NeymanConstruction: Prog: 14/100 total MC = 80 this test stat = 1.73555
s=16.2 ratioSigEff=1.00258 ratioBkgEff=1.31793 [-inf, 2.77026] in interval = 1
NeymanConstruction: Prog: 15/100 total MC = 80 this test stat = 1.64101
s=17.4 ratioSigEff=1.00283 ratioBkgEff=1.30905 [-inf, 2.84104] in interval = 1
NeymanConstruction: Prog: 16/100 total MC = 80 this test stat = 1.54913
s=18.6 ratioSigEff=1.00364 ratioBkgEff=1.29912 [-inf, 2.44366] in interval = 1
NeymanConstruction: Prog: 17/100 total MC = 80 this test stat = 1.45967
s=19.8 ratioSigEff=1.0038 ratioBkgEff=1.29047 [-inf, 2.51046] in interval = 1
NeymanConstruction: Prog: 18/100 total MC = 80 this test stat = 1.3729
s=21 ratioSigEff=1.00395 ratioBkgEff=1.2818 [-inf, 2.3088] in interval = 1
NeymanConstruction: Prog: 19/100 total MC = 80 this test stat = 1.28897
s=22.2 ratioSigEff=1.00408 ratioBkgEff=1.2731 [-inf, 2.28612] in interval = 1
NeymanConstruction: Prog: 20/100 total MC = 80 this test stat = 1.20763
s=23.4 ratioSigEff=1.0042 ratioBkgEff=1.26438 [-inf, 2.0941] in interval = 1
NeymanConstruction: Prog: 21/100 total MC = 80 this test stat = 1.12898
s=24.6 ratioSigEff=1.00431 ratioBkgEff=1.25563 [-inf, 2.07266] in interval = 1
NeymanConstruction: Prog: 22/100 total MC = 80 this test stat = 1.05305
s=25.8 ratioSigEff=1.0044 ratioBkgEff=1.24688 [-inf, 2.4819] in interval = 1
NeymanConstruction: Prog: 23/100 total MC = 80 this test stat = 0.979804
s=27 ratioSigEff=1.00448 ratioBkgEff=1.2381 [-inf, 1.94868] in interval = 1
NeymanConstruction: Prog: 24/100 total MC = 80 this test stat = 0.909232
s=28.2 ratioSigEff=1.00454 ratioBkgEff=1.22932 [-inf, 2.09127] in interval = 1
NeymanConstruction: Prog: 25/100 total MC = 80 this test stat = 0.841264
s=29.4 ratioSigEff=1.00408 ratioBkgEff=1.22097 [-inf, 1.9082] in interval = 1
NeymanConstruction: Prog: 26/100 total MC = 80 this test stat = 0.776027
s=30.6 ratioSigEff=1.00408 ratioBkgEff=1.21214 [-inf, 2.04865] in interval = 1
NeymanConstruction: Prog: 27/100 total MC = 80 this test stat = 0.713453
s=31.8 ratioSigEff=1.00406 ratioBkgEff=1.2033 [-inf, 1.4258] in interval = 1
NeymanConstruction: Prog: 28/100 total MC = 80 this test stat = 0.653398
s=33 ratioSigEff=1.00403 ratioBkgEff=1.19446 [-inf, 1.92614] in interval = 1
NeymanConstruction: Prog: 29/100 total MC = 80 this test stat = 0.596234
s=34.2 ratioSigEff=1.00399 ratioBkgEff=1.18562 [-inf, 1.53064] in interval = 1
NeymanConstruction: Prog: 30/100 total MC = 80 this test stat = 0.541678
s=35.4 ratioSigEff=1.00393 ratioBkgEff=1.17679 [-inf, 1.65725] in interval = 1
NeymanConstruction: Prog: 31/100 total MC = 80 this test stat = 0.489749
s=36.6 ratioSigEff=1.00386 ratioBkgEff=1.16795 [-inf, 1.42567] in interval = 1
NeymanConstruction: Prog: 32/100 total MC = 80 this test stat = 0.440357
s=37.8 ratioSigEff=1.00378 ratioBkgEff=1.15913 [-inf, 1.40855] in interval = 1
NeymanConstruction: Prog: 33/100 total MC = 80 this test stat = 0.393811
s=39 ratioSigEff=1.00369 ratioBkgEff=1.15032 [-inf, 1.2593] in interval = 1
NeymanConstruction: Prog: 34/100 total MC = 80 this test stat = 0.349798
s=40.2 ratioSigEff=1.00358 ratioBkgEff=1.14152 [-inf, 1.30825] in interval = 1
NeymanConstruction: Prog: 35/100 total MC = 80 this test stat = 0.30842
s=41.4 ratioSigEff=1.00347 ratioBkgEff=1.13274 [-inf, 0.874179] in interval = 1
NeymanConstruction: Prog: 36/100 total MC = 80 this test stat = 0.269673
s=42.6 ratioSigEff=1.00334 ratioBkgEff=1.12398 [-inf, 1.14949] in interval = 1
NeymanConstruction: Prog: 37/100 total MC = 80 this test stat = 0.233551
s=43.8 ratioSigEff=1.00319 ratioBkgEff=1.11573 [-inf, 1.07403] in interval = 1
NeymanConstruction: Prog: 38/100 total MC = 80 this test stat = 0.200042
s=45 ratioSigEff=1.00303 ratioBkgEff=1.10705 [-inf, 1.30848] in interval = 1
NeymanConstruction: Prog: 39/100 total MC = 80 this test stat = 0.169154
s=46.2 ratioSigEff=1.00285 ratioBkgEff=1.09838 [-inf, 0.987229] in interval = 1
NeymanConstruction: Prog: 40/100 total MC = 80 this test stat = 0.140876
s=47.4 ratioSigEff=1.00267 ratioBkgEff=1.08972 [-inf, 0.917557] in interval = 1
NeymanConstruction: Prog: 41/100 total MC = 80 this test stat = 0.115202
s=48.6 ratioSigEff=1.00247 ratioBkgEff=1.08109 [-inf, 0.850481] in interval = 1
NeymanConstruction: Prog: 42/100 total MC = 80 this test stat = 0.0921253
s=49.8 ratioSigEff=1.00226 ratioBkgEff=1.07247 [-inf, 0.891152] in interval = 1
NeymanConstruction: Prog: 43/100 total MC = 80 this test stat = 0.0716406
s=51 ratioSigEff=1.00204 ratioBkgEff=1.06387 [-inf, 1.04631] in interval = 1
NeymanConstruction: Prog: 44/100 total MC = 80 this test stat = 0.0537413
s=52.2 ratioSigEff=1.00181 ratioBkgEff=1.05529 [-inf, 0.574131] in interval = 1
NeymanConstruction: Prog: 45/100 total MC = 80 this test stat = 0.0384208
s=53.4 ratioSigEff=1.00156 ratioBkgEff=1.04672 [-inf, 0.749926] in interval = 1
NeymanConstruction: Prog: 46/100 total MC = 80 this test stat = 0.0256726
s=54.6 ratioSigEff=1.00131 ratioBkgEff=1.03818 [-inf, 0.839739] in interval = 1
NeymanConstruction: Prog: 47/100 total MC = 80 this test stat = 0.0154897
s=55.8 ratioSigEff=1.00104 ratioBkgEff=1.02966 [-inf, 0.715031] in interval = 1
NeymanConstruction: Prog: 48/100 total MC = 80 this test stat = 0.00786498
s=57 ratioSigEff=1.00075 ratioBkgEff=1.02116 [-inf, 0.574092] in interval = 1
NeymanConstruction: Prog: 49/100 total MC = 80 this test stat = 0.00303984
s=58.2 ratioSigEff=1.00144 ratioBkgEff=1.01171 [-inf, 0.731975] in interval = 1
NeymanConstruction: Prog: 50/100 total MC = 80 this test stat = 0.000296146
s=59.4 ratioSigEff=1.00049 ratioBkgEff=1.00388 [-inf, 0.692058] in interval = 1
NeymanConstruction: Prog: 51/100 total MC = 80 this test stat = 0.000277435
s=60.6 ratioSigEff=0.999518 ratioBkgEff=0.996076 [-inf, 0.645915] in interval = 1
NeymanConstruction: Prog: 52/100 total MC = 80 this test stat = 0.00297546
s=61.8 ratioSigEff=0.998533 ratioBkgEff=0.988319 [-inf, 0.513395] in interval = 1
NeymanConstruction: Prog: 53/100 total MC = 80 this test stat = 0.00784733
s=63 ratioSigEff=0.999173 ratioBkgEff=0.978981 [-inf, 0.477501] in interval = 1
NeymanConstruction: Prog: 54/100 total MC = 80 this test stat = 0.0154172
s=64.2 ratioSigEff=0.998823 ratioBkgEff=0.970615 [-inf, 0.721422] in interval = 1
NeymanConstruction: Prog: 55/100 total MC = 80 this test stat = 0.0254769
s=65.4 ratioSigEff=0.998461 ratioBkgEff=0.962272 [-inf, 0.678264] in interval = 1
NeymanConstruction: Prog: 56/100 total MC = 80 this test stat = 0.0380399
s=66.6 ratioSigEff=0.998088 ratioBkgEff=0.953953 [-inf, 0.636603] in interval = 1
NeymanConstruction: Prog: 57/100 total MC = 80 this test stat = 0.0530874
s=67.8 ratioSigEff=0.997704 ratioBkgEff=0.945659 [-inf, 0.596425] in interval = 1
NeymanConstruction: Prog: 58/100 total MC = 80 this test stat = 0.0706058
s=69 ratioSigEff=0.997308 ratioBkgEff=0.93739 [-inf, 0.70275] in interval = 1
NeymanConstruction: Prog: 59/100 total MC = 80 this test stat = 0.0905894
s=70.2 ratioSigEff=0.996902 ratioBkgEff=0.929146 [-inf, 0.93255] in interval = 1
NeymanConstruction: Prog: 60/100 total MC = 80 this test stat = 0.11303
s=71.4 ratioSigEff=0.996485 ratioBkgEff=0.920929 [-inf, 1.06482] in interval = 1
NeymanConstruction: Prog: 61/100 total MC = 80 this test stat = 0.137918
s=72.6 ratioSigEff=0.996058 ratioBkgEff=0.912737 [-inf, 0.780337] in interval = 1
NeymanConstruction: Prog: 62/100 total MC = 80 this test stat = 0.165246
s=73.8 ratioSigEff=0.995619 ratioBkgEff=0.904572 [-inf, 0.684448] in interval = 1
NeymanConstruction: Prog: 63/100 total MC = 80 this test stat = 0.195001
s=75 ratioSigEff=0.99517 ratioBkgEff=0.896433 [-inf, 1.03027] in interval = 1
NeymanConstruction: Prog: 64/100 total MC = 80 this test stat = 0.227176
s=76.2 ratioSigEff=0.99471 ratioBkgEff=0.888322 [-inf, 0.978263] in interval = 1
NeymanConstruction: Prog: 65/100 total MC = 80 this test stat = 0.261776
s=77.4 ratioSigEff=0.99424 ratioBkgEff=0.880239 [-inf, 1.11256] in interval = 1
NeymanConstruction: Prog: 66/100 total MC = 80 this test stat = 0.298772
s=78.6 ratioSigEff=0.99376 ratioBkgEff=0.872183 [-inf, 1.18776] in interval = 1
NeymanConstruction: Prog: 67/100 total MC = 80 this test stat = 0.338144
s=79.8 ratioSigEff=0.993269 ratioBkgEff=0.864156 [-inf, 1.48101] in interval = 1
NeymanConstruction: Prog: 68/100 total MC = 80 this test stat = 0.379914
s=81 ratioSigEff=0.992769 ratioBkgEff=0.856157 [-inf, 1.56755] in interval = 1
NeymanConstruction: Prog: 69/100 total MC = 80 this test stat = 0.424071
s=82.2 ratioSigEff=0.992258 ratioBkgEff=0.848187 [-inf, 1.21708] in interval = 1
NeymanConstruction: Prog: 70/100 total MC = 80 this test stat = 0.470561
s=83.4 ratioSigEff=0.991737 ratioBkgEff=0.840247 [-inf, 1.43809] in interval = 1
NeymanConstruction: Prog: 71/100 total MC = 80 this test stat = 0.519388
s=84.6 ratioSigEff=0.991206 ratioBkgEff=0.832336 [-inf, 1.84094] in interval = 1
NeymanConstruction: Prog: 72/100 total MC = 80 this test stat = 0.570571
s=85.8 ratioSigEff=0.990665 ratioBkgEff=0.824455 [-inf, 1.31514] in interval = 1
NeymanConstruction: Prog: 73/100 total MC = 80 this test stat = 0.624119
s=87 ratioSigEff=0.990114 ratioBkgEff=0.816605 [-inf, 1.18918] in interval = 1
NeymanConstruction: Prog: 74/100 total MC = 80 this test stat = 0.679954
s=88.2 ratioSigEff=0.989554 ratioBkgEff=0.808785 [-inf, 1.33503] in interval = 1
NeymanConstruction: Prog: 75/100 total MC = 80 this test stat = 0.738105
s=89.4 ratioSigEff=0.988984 ratioBkgEff=0.800996 [-inf, 1.56456] in interval = 1
NeymanConstruction: Prog: 76/100 total MC = 80 this test stat = 0.798548
s=90.6 ratioSigEff=0.988404 ratioBkgEff=0.793239 [-inf, 1.49965] in interval = 1
NeymanConstruction: Prog: 77/100 total MC = 80 this test stat = 0.861306
s=91.8 ratioSigEff=0.987815 ratioBkgEff=0.785512 [-inf, 1.66275] in interval = 1
NeymanConstruction: Prog: 78/100 total MC = 80 this test stat = 0.926319
s=93 ratioSigEff=0.987217 ratioBkgEff=0.777818 [-inf, 1.44643] in interval = 1
NeymanConstruction: Prog: 79/100 total MC = 80 this test stat = 0.993583
s=94.2 ratioSigEff=0.98661 ratioBkgEff=0.770156 [-inf, 2.19051] in interval = 1
NeymanConstruction: Prog: 80/100 total MC = 80 this test stat = 1.06308
s=95.4 ratioSigEff=0.985993 ratioBkgEff=0.762527 [-inf, 1.54054] in interval = 1
NeymanConstruction: Prog: 81/100 total MC = 80 this test stat = 1.13458
s=96.6 ratioSigEff=0.985367 ratioBkgEff=0.75493 [-inf, 1.86615] in interval = 1
NeymanConstruction: Prog: 82/100 total MC = 80 this test stat = 1.20859
s=97.8 ratioSigEff=0.984732 ratioBkgEff=0.747367 [-inf, 2.40993] in interval = 1
NeymanConstruction: Prog: 83/100 total MC = 80 this test stat = 1.28501
s=99 ratioSigEff=0.984088 ratioBkgEff=0.739836 [-inf, 2.32724] in interval = 1
NeymanConstruction: Prog: 84/100 total MC = 80 this test stat = 1.36342
s=100.2 ratioSigEff=0.983435 ratioBkgEff=0.73234 [-inf, 2.43281] in interval = 1
NeymanConstruction: Prog: 85/100 total MC = 80 this test stat = 1.44405
s=101.4 ratioSigEff=0.982773 ratioBkgEff=0.724877 [-inf, 2.44403] in interval = 1
NeymanConstruction: Prog: 86/100 total MC = 80 this test stat = 1.52687
s=102.6 ratioSigEff=0.982103 ratioBkgEff=0.717449 [-inf, 2.09032] in interval = 1
NeymanConstruction: Prog: 87/100 total MC = 80 this test stat = 1.61182
s=103.8 ratioSigEff=0.981424 ratioBkgEff=0.710056 [-inf, 2.76135] in interval = 1
NeymanConstruction: Prog: 88/100 total MC = 80 this test stat = 1.69896
s=105 ratioSigEff=0.980736 ratioBkgEff=0.702697 [-inf, 2.38331] in interval = 1
NeymanConstruction: Prog: 89/100 total MC = 80 this test stat = 1.78823
s=106.2 ratioSigEff=0.98004 ratioBkgEff=0.695373 [-inf, 2.39444] in interval = 1
NeymanConstruction: Prog: 90/100 total MC = 80 this test stat = 1.87929
s=107.4 ratioSigEff=0.979335 ratioBkgEff=0.688084 [-inf, 2.89798] in interval = 1
NeymanConstruction: Prog: 91/100 total MC = 80 this test stat = 1.97313
s=108.6 ratioSigEff=0.978622 ratioBkgEff=0.680832 [-inf, 3.1204] in interval = 1
NeymanConstruction: Prog: 92/100 total MC = 80 this test stat = 2.06825
s=109.8 ratioSigEff=0.977383 ratioBkgEff=0.670641 [-inf, 3.13169] in interval = 1
NeymanConstruction: Prog: 93/100 total MC = 80 this test stat = 2.16592
s=111 ratioSigEff=0.976625 ratioBkgEff=0.663286 [-inf, 2.93195] in interval = 1
NeymanConstruction: Prog: 94/100 total MC = 80 this test stat = 2.26567
s=112.2 ratioSigEff=0.975857 ratioBkgEff=0.655962 [-inf, 2.54424] in interval = 1
NeymanConstruction: Prog: 95/100 total MC = 80 this test stat = 2.36747
s=113.4 ratioSigEff=0.975049 ratioBkgEff=0.648489 [-inf, 3.05911] in interval = 1
NeymanConstruction: Prog: 96/100 total MC = 80 this test stat = 2.47112
s=114.6 ratioSigEff=0.97426 ratioBkgEff=0.641206 [-inf, 2.96555] in interval = 1
NeymanConstruction: Prog: 97/100 total MC = 80 this test stat = 2.57725
s=115.8 ratioSigEff=0.973462 ratioBkgEff=0.633952 [-inf, 3.5205] in interval = 1
NeymanConstruction: Prog: 98/100 total MC = 240 this test stat = 2.68512
s=117 ratioSigEff=0.972655 ratioBkgEff=0.626728 [-inf, 3.19934] in interval = 1
NeymanConstruction: Prog: 99/100 total MC = 80 this test stat = 2.79507
s=118.2 ratioSigEff=0.971839 ratioBkgEff=0.619534 [-inf, 3.77433] in interval = 1
NeymanConstruction: Prog: 100/100 total MC = 80 this test stat = 2.90706
s=119.4 ratioSigEff=0.971013 ratioBkgEff=0.61237 [-inf, 3.66884] in interval = 1
[#1] INFO:Eval -- 100 points in interval
[#1] INFO:Minimization -- Including the following constraint terms in minimization: (sigConstraint,bkgConstraint)
[#1] INFO:Minimization -- The global observables are not defined , normalize constraints with respect to the parameters (b,ratioBkgEff,ratioSigEff,s)
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_modelWithConstraints_exampleData_with_constr) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- The following expressions will be evaluated in cache-and-track mode: (countingModel)
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
[#1] INFO:Minimization -- Including the following constraint terms in minimization: (sigConstraint,bkgConstraint)
[#1] INFO:Minimization -- The following global observables have been defined and their values are taken from the model: (gSigEff,gSigBkg)
Metropolis-Hastings progress: ....................................................................................................
[#1] INFO:Eval -- Proposal acceptance rate: 48.165%
[#1] INFO:Eval -- Number of steps in chain: 9633
Real time 0:00:12, CP time 12.410
Profile lower limit on s = 13.9497380406319
Profile upper limit on s = 107.9503617088649
FC lower limit on s = 0.6
FC upper limit on s = 119.39999999999999
MCMC lower limit on s = 19.470961227250626
MCMC upper limit on s = 104.4475703575416
MCMC Actual confidence level: 0.9500150557061127
plotting the chain data - nentries = 9633
plotting the scanned points used in the frequentist construction - npoints = 100
import ROOT
t = ROOT.TStopwatch()
t.Start()
wspace = ROOT.RooWorkspace()
wspace.factory(
"Poisson::countingModel(obs[150,0,300], " "sum(s[50,0,120]*ratioSigEff[1.,0,3.],b[100]*ratioBkgEff[1.,0.,3.]))"
)
wspace.factory("Gaussian::sigConstraint(gSigEff[1,0,3],ratioSigEff,0.05)")
wspace.factory("Gaussian::bkgConstraint(gSigBkg[1,0,3],ratioBkgEff,0.2)")
wspace.factory("PROD::modelWithConstraints(countingModel,sigConstraint,bkgConstraint)")
wspace.Print()
modelWithConstraints = wspace["modelWithConstraints"]
obs = wspace["obs"]
s = wspace["s"]
b = wspace["b"]
b.setConstant()
ratioSigEff = wspace["ratioSigEff"]
ratioBkgEff = wspace["ratioBkgEff"]
constrainedParams = {ratioSigEff, ratioBkgEff}
gSigEff = wspace["gSigEff"]
gSigBkg = wspace["gSigBkg"]
gSigEff.setConstant()
gSigBkg.setConstant()
obs.setVal(160.0)
data = ROOT.RooDataSet("exampleData", "exampleData", {obs})
data.add(obs)
modelWithConstraints.fitTo(data, Constrain=constrainedParams, PrintLevel=-1)
modelConfig = ROOT.RooStats.ModelConfig(wspace)
modelConfig.SetPdf(modelWithConstraints)
modelConfig.SetParametersOfInterest({s})
modelConfig.SetNuisanceParameters(constrainedParams)
modelConfig.SetObservables(obs)
modelConfig.SetGlobalObservables({gSigEff, gSigBkg})
modelConfig.SetName("ModelConfig")
wspace.Import(modelConfig)
wspace.Import(data)
wspace.SetName("w")
wspace.writeToFile("rs101_ws.root")
plc = ROOT.RooStats.ProfileLikelihoodCalculator(data, modelConfig)
plc.SetTestSize(0.05)
lrinterval = plc.GetInterval()
dataCanvas = ROOT.TCanvas("dataCanvas")
dataCanvas.Divide(2, 1)
dataCanvas.cd(1)
plotInt = ROOT.RooStats.LikelihoodIntervalPlot(lrinterval)
plotInt.SetTitle("Profile Likelihood Ratio and Posterior for S")
plotInt.Draw()
fc = ROOT.RooStats.FeldmanCousins(data, modelConfig)
fc.UseAdaptiveSampling(True)
fc.FluctuateNumDataEntries(False)
fc.SetNBins(100)
fc.SetTestSize(0.05)
fcint = fc.GetInterval()
fit = modelWithConstraints.fitTo(data, Save=True, PrintLevel=-1)
ph = ROOT.RooStats.ProposalHelper()
ph.SetVariables(fit.floatParsFinal())
ph.SetCovMatrix(fit.covarianceMatrix())
ph.SetUpdateProposalParameters(True)
ph.SetCacheSize(100)
pdfProp = ph.GetProposalFunction()
mc = ROOT.RooStats.MCMCCalculator(data, modelConfig)
mc.SetNumIters(20000)
mc.SetTestSize(0.05)
mc.SetNumBurnInSteps(40)
mc.SetProposalFunction(pdfProp)
mc.SetLeftSideTailFraction(0.5)
mcInt = mc.GetInterval()
print("Profile lower limit on s = ", lrinterval.LowerLimit(s))
print("Profile upper limit on s = ", lrinterval.UpperLimit(s))
if fcint:
fcul = fcint.UpperLimit(s)
fcll = fcint.LowerLimit(s)
print("FC lower limit on s = ", fcll)
print("FC upper limit on s = ", fcul)
fcllLine = ROOT.TLine(fcll, 0, fcll, 1)
fculLine = ROOT.TLine(fcul, 0, fcul, 1)
fcllLine.SetLineColor(ROOT.kRed)
fculLine.SetLineColor(ROOT.kRed)
fcllLine.Draw("same")
fculLine.Draw("same")
dataCanvas.Update()
mcPlot = ROOT.RooStats.MCMCIntervalPlot(mcInt)
mcPlot.SetLineColor(ROOT.kMagenta)
mcPlot.SetLineWidth(2)
mcPlot.Draw("same")
mcul = mcInt.UpperLimit(s)
mcll = mcInt.LowerLimit(s)
print("MCMC lower limit on s = ", mcll)
print("MCMC upper limit on s = ", mcul)
print("MCMC Actual confidence level: ", mcInt.GetActualConfidenceLevel())
dataCanvas.cd(2)
chainData = mcInt.GetChainAsDataSet()
print("plotting the chain data - nentries = ", chainData.numEntries())
chain = ROOT.RooStats.GetAsTTree("chainTreeData", "chainTreeData", chainData)
chain.SetMarkerStyle(6)
chain.SetMarkerColor(ROOT.kRed)
chain.Draw("s:ratioSigEff:ratioBkgEff", "nll_MarkovChain_local_", "box")
parScanData = fc.GetPointsToScan()
print("plotting the scanned points used in the frequentist construction - npoints = ", parScanData.numEntries())
gr = ROOT.TGraph2D(parScanData.numEntries())
for ievt in range(parScanData.numEntries()):
evt = parScanData.get(ievt)
x = evt.getRealValue("ratioBkgEff")
y = evt.getRealValue("ratioSigEff")
z = evt.getRealValue("s")
gr.SetPoint(ievt, x, y, z)
gr.SetMarkerStyle(24)
gr.Draw("P SAME")
t.Stop()
t.Print()
dataCanvas.SaveAs("rs101_limitexample.png")
del mc