With default parameters the macro will attempt to run the standard hist2workspace example and read the ROOT file that it produces.
The actual heart of the demo is only about 10 lines long.
The FeldmanCousins tools is a classical frequentist calculation based on the Neyman Construction. The test statistic can be generalized for nuisance parameters by using the profile likelihood ratio. But unlike the ProfileLikelihoodCalculator, this tool explicitly builds the sampling distribution of the test statistic via toy Monte Carlo.
=== Using the following for ModelConfig ===
Observables: RooArgSet:: = (obs_x_channel1,channelCat)
Parameters of Interest: RooArgSet:: = (SigXsecOverSM)
Nuisance Parameters: RooArgSet:: = (alpha_syst2,alpha_syst3,gamma_stat_channel1_bin_0,gamma_stat_channel1_bin_1)
Global Observables: RooArgSet:: = (nominalLumi,nom_alpha_syst1,nom_alpha_syst2,nom_alpha_syst3,nom_gamma_stat_channel1_bin_0,nom_gamma_stat_channel1_bin_1)
PDF: RooSimultaneous::simPdf[ indexCat=channelCat channel1=model_channel1 ] = 0.190787
FeldmanCousins: ntoys per point: adaptive
FeldmanCousins: nEvents per toy will fluctuate about expectation
FeldmanCousins: Model has nuisance parameters, will do profile construction
FeldmanCousins: # points to test = 10
lookup index = 0
NeymanConstruction: Prog: 1/10 total MC = 78 this test stat = 1.32297
SigXsecOverSM=0.15 alpha_syst2=0.602143 alpha_syst3=0.227772 gamma_stat_channel1_bin_0=1.03121 gamma_stat_channel1_bin_1=1.04626 [-inf, 3.00302] in interval = 1
NeymanConstruction: Prog: 2/10 total MC = 78 this test stat = 0.622271
SigXsecOverSM=0.45 alpha_syst2=0.407504 alpha_syst3=0.166261 gamma_stat_channel1_bin_0=1.0207 gamma_stat_channel1_bin_1=1.03105 [-inf, 2.56102] in interval = 1
NeymanConstruction: Prog: 3/10 total MC = 78 this test stat = 0.185599
SigXsecOverSM=0.75 alpha_syst2=0.213102 alpha_syst3=0.0987913 gamma_stat_channel1_bin_0=1.01077 gamma_stat_channel1_bin_1=1.01853 [-inf, 1.34636] in interval = 1
NeymanConstruction: Prog: 4/10 total MC = 78 this test stat = 0.00589021
SigXsecOverSM=1.05 alpha_syst2=0.0285809 alpha_syst3=0.0409103 gamma_stat_channel1_bin_0=1.00142 gamma_stat_channel1_bin_1=1.00599 [-inf, 1.11873] in interval = 1
NeymanConstruction: Prog: 5/10 total MC = 78 this test stat = 0.0746369
SigXsecOverSM=1.35 alpha_syst2=-0.137558 alpha_syst3=-0.0496314 gamma_stat_channel1_bin_0=0.993073 gamma_stat_channel1_bin_1=0.994154 [-inf, 2.18034] in interval = 1
NeymanConstruction: Prog: 6/10 total MC = 78 this test stat = 0.383697
SigXsecOverSM=1.65 alpha_syst2=-0.307933 alpha_syst3=-0.098712 gamma_stat_channel1_bin_0=0.984872 gamma_stat_channel1_bin_1=0.982202 [-inf, 1.23693] in interval = 1
NeymanConstruction: Prog: 7/10 total MC = 78 this test stat = 0.924084
SigXsecOverSM=1.95 alpha_syst2=-0.46183 alpha_syst3=-0.162017 gamma_stat_channel1_bin_0=0.977442 gamma_stat_channel1_bin_1=0.970551 [-inf, 1.89854] in interval = 1
NeymanConstruction: Prog: 8/10 total MC = 78 this test stat = 1.68896
SigXsecOverSM=2.25 alpha_syst2=-0.604401 alpha_syst3=-0.223564 gamma_stat_channel1_bin_0=0.970553 gamma_stat_channel1_bin_1=0.959142 [-inf, 2.02832] in interval = 1
NeymanConstruction: Prog: 9/10 total MC = 234 this test stat = 2.66964
SigXsecOverSM=2.55 alpha_syst2=-0.735442 alpha_syst3=-0.283315 gamma_stat_channel1_bin_0=0.964167 gamma_stat_channel1_bin_1=0.947986 [-inf, 2.14772] in interval = 0
NeymanConstruction: Prog: 10/10 total MC = 234 this test stat = 3.85849
SigXsecOverSM=2.85 alpha_syst2=-0.855772 alpha_syst3=-0.341177 gamma_stat_channel1_bin_0=0.958246 gamma_stat_channel1_bin_1=0.937094 [-inf, 2.28493] in interval = 0
[#1] INFO:Eval -- 8 points in interval
95% interval on SigXsecOverSM is : [0.15, 2.25]
void StandardFeldmanCousinsDemo(const char *infile = "", const char *workspaceName = "combined",
const char *modelConfigName = "ModelConfig", const char *dataName = "obsData")
{
if (!strcmp(infile, "")) {
filename =
"results/example_combined_GaussExample_model.root";
if (!fileExist) {
cout << "will run standard hist2workspace example" << endl;
gROOT->ProcessLine(
".! prepareHistFactory .");
gROOT->ProcessLine(
".! hist2workspace config/example.xml");
cout << "\n\n---------------------" << endl;
cout << "Done creating example input" << endl;
cout << "---------------------\n\n" << endl;
}
} else
cout <<
"StandardRooStatsDemoMacro: Input file " <<
filename <<
" is not found" << endl;
return;
}
cout << "workspace not found" << endl;
return;
}
cout << "data or ModelConfig was not found" << endl;
return;
}
fc.SetConfidenceLevel(0.95);
fc.UseAdaptiveSampling(true);
fc.SetNBins(10);
fc.CreateConfBelt(true);
if (
data->numEntries() == 1)
fc.FluctuateNumDataEntries(false);
else
cout << "Not sure what to do about this model" << endl;
}
cout <<
"\n95% interval on " << firstPOI->
GetName() <<
" is : [" << interval->
LowerLimit(*firstPOI) <<
", " << interval->
UpperLimit(*firstPOI) <<
"] " << endl;
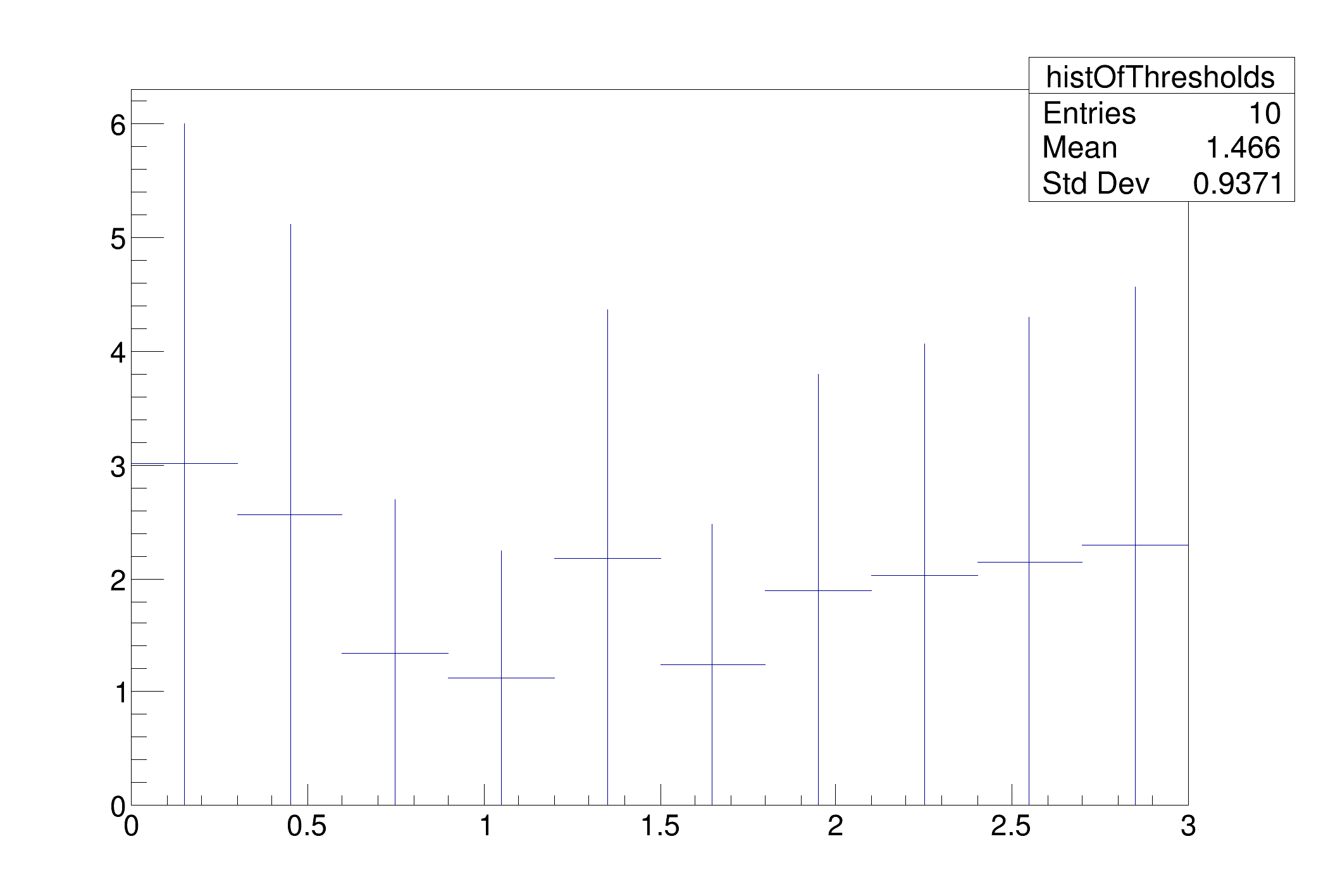
histOfThresholds->
Fill(poiVal, arMax);
}
histOfThresholds->
Draw();
}
Option_t Option_t TPoint TPoint const char GetTextMagnitude GetFillStyle GetLineColor GetLineWidth GetMarkerStyle GetTextAlign GetTextColor GetTextSize void data
Option_t Option_t TPoint TPoint const char GetTextMagnitude GetFillStyle GetLineColor GetLineWidth GetMarkerStyle GetTextAlign GetTextColor GetTextSize void char Point_t Rectangle_t WindowAttributes_t Float_t Float_t Float_t Int_t Int_t UInt_t UInt_t Rectangle_t Int_t Int_t Window_t TString Int_t GCValues_t GetPrimarySelectionOwner GetDisplay GetScreen GetColormap GetNativeEvent const char const char dpyName wid window const char font_name cursor keysym reg const char only_if_exist regb h Point_t winding char text const char depth char const char Int_t count const char ColorStruct_t color const char filename
R__EXTERN TSystem * gSystem
double getRealValue(const char *name, double defVal=0.0, bool verbose=false) const
Get value of a RooAbsReal stored in set with given name.
RooAbsArg * first() const
Abstract base class for binned and unbinned datasets.
virtual Int_t numEntries() const
Return number of entries in dataset, i.e., count unweighted entries.
bool canBeExtended() const
If true, PDF can provide extended likelihood term.
virtual double getMax(const char *name=nullptr) const
Get maximum of currently defined range.
virtual double getMin(const char *name=nullptr) const
Get minimum of currently defined range.
RooArgSet is a container object that can hold multiple RooAbsArg objects.
TObject * clone(const char *newname) const override
RooDataSet is a container class to hold unbinned data.
const RooArgSet * get(Int_t index) const override
Return RooArgSet with coordinates of event 'index'.
RooRealVar represents a variable that can be changed from the outside.
ConfidenceBelt is a concrete implementation of the ConfInterval interface.
double GetAcceptanceRegionMax(RooArgSet &, double cl=-1., double leftside=-1.)
The FeldmanCousins class (like the Feldman-Cousins technique) is essentially a specific configuration...
ModelConfig is a simple class that holds configuration information specifying how a model should be u...
const RooArgSet * GetParametersOfInterest() const
get RooArgSet containing the parameter of interest (return nullptr if not existing)
RooAbsPdf * GetPdf() const
get model PDF (return nullptr if pdf has not been specified or does not exist)
PointSetInterval is a concrete implementation of the ConfInterval interface.
double UpperLimit(RooRealVar ¶m)
return upper limit on a given parameter
double LowerLimit(RooRealVar ¶m)
return lower limit on a given parameter
Persistable container for RooFit projects.
A ROOT file is composed of a header, followed by consecutive data records (TKey instances) with a wel...
static TFile * Open(const char *name, Option_t *option="", const char *ftitle="", Int_t compress=ROOT::RCompressionSetting::EDefaults::kUseCompiledDefault, Int_t netopt=0)
Create / open a file.
1-D histogram with a float per channel (see TH1 documentation)}
virtual Int_t Fill(Double_t x)
Increment bin with abscissa X by 1.
void Draw(Option_t *option="") override
Draw this histogram with options.
virtual void SetMinimum(Double_t minimum=-1111)
const char * GetName() const override
Returns name of object.
virtual Bool_t AccessPathName(const char *path, EAccessMode mode=kFileExists)
Returns FALSE if one can access a file using the specified access mode.
The namespace RooFit contains mostly switches that change the behaviour of functions of PDFs (or othe...
Namespace for the RooStats classes.