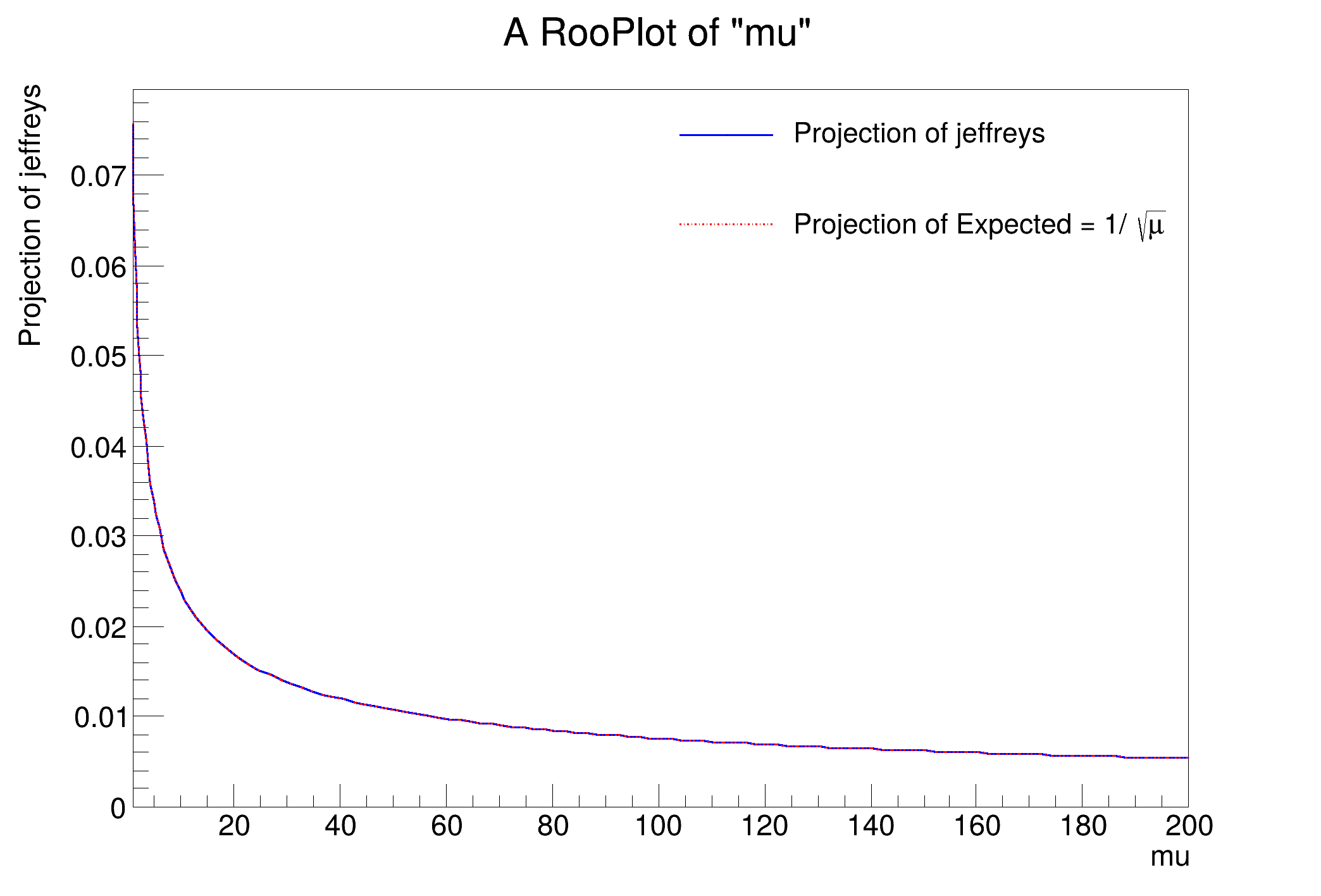
Jeffreys's prior is an 'objective prior' based on formal rules. It is calculated from the Fisher information matrix.
The analytic form is not known for most PDFs, but it is for simple cases like the Poisson mean, Gaussian mean, Gaussian sigma.
This class uses numerical tricks to calculate the Fisher Information Matrix efficiently. In particular, it takes advantage of a property of the 'Asimov data' as described in Asymptotic formulae for likelihood-based tests of new physics Glen Cowan, Kyle Cranmer, Eilam Gross, Ofer Vitells http://arxiv.org/abs/arXiv:1007.1727
[#1] INFO:Minimization -- p.d.f. provides expected number of events, including extended term in likelihood.
[#1] INFO:Fitting -- RooAbsPdf::fitTo(p) fixing normalization set for coefficient determination to observables in data
[#1] INFO:Fitting -- using generic CPU library compiled with no vectorizations
[#1] INFO:Fitting -- Creation of NLL object took 794.154 μs
[#1] INFO:Fitting -- RooAddition::defaultErrorLevel(nll_p_genData) Summation contains a RooNLLVar, using its error level
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: activating const optimization
[#1] INFO:Minimization -- [fitFCN] No discrete parameters, performing continuous minimization only
Minuit2Minimizer: Minimize with max-calls 500 convergence for edm < 1 strategy 1
Minuit2Minimizer : Valid minimum - status = 0
FVAL = -360.517018598809159
Edm = 4.13877261906962124e-17
Nfcn = 22
mu = 100 +/- 9.98317 (limited)
[#1] INFO:Fitting -- RooAbsPdf::fitTo(p) Calculating sum-of-weights-squared correction matrix for covariance matrix
[#1] INFO:Minimization -- RooAbsMinimizerFcn::setOptimizeConst: deactivating const optimization
RooDataHist::genData[x] = 100 bins (100 weights)
RooFitResult: minimized FCN value: -360.517, estimated distance to minimum: 1.4664e-19
covariance matrix quality: Full, accurate covariance matrix
Status : MINIMIZE=0 HESSE=0 HESSE=0
Floating Parameter FinalValue +/- Error
-------------------- --------------------------
mu 1.0000e+02 +/- 1.00e+01
variance = 100
stdev = 10
jeffreys = 0.0999998
[#1] INFO:NumericIntegration -- RooRealIntegral::init(jeffreys_Int[mu]) using numeric integrator RooAdaptiveGaussKronrodIntegrator1D to calculate Int(mu)
[#1] INFO:NumericIntegration -- RooRealIntegral::init(Expected_Int[mu]) using numeric integrator RooIntegrator1D to calculate Int(mu)
void rs302_JeffreysPriorDemo()
{
w.factory("Uniform::u(x[0,1])");
w.factory("mu[100,1,200]");
w.factory("ExtendPdf::p(u,mu)");
std::unique_ptr<RooDataHist> asimov{w.pdf(
"p")->generateBinned(*w.var(
"x"),
ExpectedData())};
std::unique_ptr<RooFitResult> res{w.pdf(
"p")->fitTo(*asimov,
Save(),
SumW2Error(
true))};
asimov->Print();
res->Print();
w.defineSet("poi", "mu");
w.defineSet("obs", "x");
*w.set("poi"));
RooPlot *plot = w.var(
"mu")->frame();
pi.plotOn(plot);
legend->DrawClone();
}
void TestJeffreysGaussMean()
{
w.factory("Gaussian::g(x[0,-20,20],mu[0,-5.,5],sigma[1,0,10])");
w.factory("n[10,.1,200]");
w.factory("ExtendPdf::p(g,n)");
w.var("sigma")->setConstant();
w.var("n")->setConstant();
std::unique_ptr<RooDataHist> asimov{w.pdf(
"p")->generateBinned(*w.var(
"x"),
ExpectedData())};
asimov->Print();
res->Print();
w.defineSet("poi", "mu");
w.defineSet("obs", "x");
pi.getParameters(*temp)->Print();
RooPlot *plot = w.var(
"mu")->frame();
pi.plotOn(plot);
legend->DrawClone();
}
void TestJeffreysGaussSigma()
{
w.factory("Gaussian::g(x[0,-20,20],mu[0,-5,5],sigma[1,1,5])");
w.factory("n[100,.1,2000]");
w.factory("ExtendPdf::p(g,n)");
w.var("mu")->setConstant();
w.var("n")->setConstant();
w.var("x")->setBins(301);
std::unique_ptr<RooDataHist> asimov{w.pdf(
"p")->generateBinned(*w.var(
"x"),
ExpectedData())};
asimov->Print();
res->Print();
w.defineSet("poi", "sigma");
w.defineSet("obs", "x");
pi.getParameters(*temp)->Print();
RooPlot *plot = w.var(
"sigma")->frame();
pi.plotOn(plot);
legend->DrawClone();
}
void TestJeffreysGaussMeanAndSigma()
{
w.factory("Gaussian::g(x[0,-20,20],mu[0,-5,5],sigma[1,1.,5.])");
w.factory("n[100,.1,2000]");
w.factory("ExtendPdf::p(g,n)");
w.var("n")->setConstant();
w.var("x")->setBins(301);
std::unique_ptr<RooDataHist> asimov{w.pdf(
"p")->generateBinned(*w.var(
"x"),
ExpectedData())};
asimov->Print();
res->Print();
w.defineSet("poi", "mu,sigma");
w.defineSet("obs", "x");
pi.getParameters(*temp)->Print();
}
TMatrixTSym< Double_t > TMatrixDSym
RooPlot * plotOn(RooPlot *frame, const RooCmdArg &arg1={}, const RooCmdArg &arg2={}, const RooCmdArg &arg3={}, const RooCmdArg &arg4={}, const RooCmdArg &arg5={}, const RooCmdArg &arg6={}, const RooCmdArg &arg7={}, const RooCmdArg &arg8={}, const RooCmdArg &arg9={}, const RooCmdArg &arg10={}) const override
Helper calling plotOn(RooPlot*, RooLinkedList&) const.
RooArgSet is a container object that can hold multiple RooAbsArg objects.
Implementation of a probability density function that takes a RooArgList of servers and a C++ express...
Implementation of Jeffrey's prior.
Plot frame and a container for graphics objects within that frame.
void Draw(Option_t *options=nullptr) override
Draw this plot and all of the elements it contains.
std::unique_ptr< TLegend > BuildLegend() const
Build a legend that contains all objects that have been drawn on the plot.
Persistable container for RooFit projects.
TH1 is the base class of all histogram classes in ROOT.
void Draw(Option_t *option="") override
Draw this histogram with options.
Double_t Determinant() const override
TMatrixTSym< Element > & Invert(Double_t *det=nullptr)
Invert the matrix and calculate its determinant Notice that the LU decomposition is used instead of B...
RooCmdArg YVar(const RooAbsRealLValue &var, const RooCmdArg &arg={})
RooCmdArg Save(bool flag=true)
RooCmdArg SumW2Error(bool flag)
RooCmdArg ExpectedData(bool flag=true)
RooCmdArg Binning(const RooAbsBinning &binning)
RooCmdArg LineColor(TColorNumber color)
RooCmdArg LineStyle(Style_t style)
VecExpr< UnaryOp< Sqrt< T >, VecExpr< A, T, D >, T >, T, D > sqrt(const VecExpr< A, T, D > &rhs)
The namespace RooFit contains mostly switches that change the behaviour of functions of PDFs (or othe...