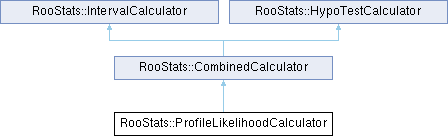
The ProfileLikelihoodCalculator is a concrete implementation of CombinedCalculator (the interface class for tools which can produce both a RooStats HypoTestResult and ConfInterval).
The tool uses the profile likelihood ratio as a test statistic, and assumes that Wilks' theorem is valid. Wilks' theorem states that \( -2 \cdot \ln(\lambda) \) (profile likelihood ratio) is asymptotically distributed as a \( \chi^2 \) distribution with \( N \) degrees of freedom. Thus, \(p\)-values can be constructed, and the profile likelihood ratio can be used to construct a LikelihoodInterval. (In the future, this class could be extended to use toy Monte Carlo to calibrate the distribution of the test statistic).
Usage: It uses the interface of the CombinedCalculator, so that it can be configured by specifying:
The interface allows one to pass the model, data, and parameters either directly or via a ModelConfig class. The alternate hypothesis leaves the parameter free to take any value other than those specified by the null hypothesis. There is therefore no need to specify the alternate parameters.
After configuring the calculator, one only needs to call GetHypoTest() (which will return a HypoTestResult pointer) or GetInterval() (which will return a ConfInterval pointer).
This calculator can work with both one-dimensional intervals or multi- dimensional ones (contours).
Note that for hypothesis tests, it is often better to use the AsymptoticCalculator, which can compute in addition the expected \(p\)-value using an Asimov data set.
Definition at line 22 of file ProfileLikelihoodCalculator.h.
Public Member Functions | |
| ProfileLikelihoodCalculator () | |
| Default constructor (needed for I/O). | |
| ProfileLikelihoodCalculator (RooAbsData &data, ModelConfig &model, double size=0.05) | |
| Constructor from data and a model configuration If the ModelConfig defines a prior pdf for any of the parameters those will be included as constrained terms in the likelihood function. | |
| ProfileLikelihoodCalculator (RooAbsData &data, RooAbsPdf &pdf, const RooArgSet ¶msOfInterest, double size=0.05, const RooArgSet *nullParams=nullptr) | |
| Constructor from data, from a full model pdf describing both parameter of interest and nuisance parameters and from the set specifying the parameter of interest (POI). | |
| ~ProfileLikelihoodCalculator () override | |
| destructor cannot delete prod pdf because it will delete all the composing pdf's if (fOwnPdf) delete fPdf; fPdf = 0; | |
| double | ConfidenceLevel () const override |
| Get the Confidence level for the test. | |
| HypoTestResult * | GetHypoTest () const override |
| Return the hypothesis test result obtained from the likelihood ratio of the maximum likelihood value with the null parameters fixed to their values, with respect to keeping all parameters floating (global maximum likelihood value). | |
| LikelihoodInterval * | GetInterval () const override |
| Return a likelihood interval. | |
| TClass * | IsA () const override |
| void | SetAlternateModel (const ModelConfig &) override |
| Set the model for the alternate hypothesis. | |
| virtual void | SetAlternateParameters (const RooArgSet &set) |
| set parameter values for the alternate if using a common PDF | |
| virtual void | SetCommonModel (const ModelConfig &model) |
| Set a common model for both the null and alternate, add to the workspace if not already there. | |
| virtual void | SetConditionalObservables (const RooArgSet &set) |
| set conditional observables needed for computing the NLL | |
| void | SetConfidenceLevel (double cl) override |
| set the confidence level for the interval (eg. 0.95 for a 95% Confidence Interval) | |
| void | SetData (RooAbsData &data) override |
| Set the DataSet, add to the workspace if not already there. | |
| virtual void | SetGlobalObservables (const RooArgSet &set) |
| set global observables needed for computing the NLL | |
| void | SetModel (const ModelConfig &model) override |
| set the model (in this case can set only the parameters for the null hypothesis) | |
| virtual void | SetNuisanceParameters (const RooArgSet &set) |
| specify the nuisance parameters (eg. the rest of the parameters) | |
| void | SetNullModel (const ModelConfig &) override |
| Set the model for the null hypothesis. | |
| virtual void | SetNullParameters (const RooArgSet &set) |
| set parameter values for the null if using a common PDF | |
| virtual void | SetParameters (const RooArgSet &set) |
| specify the parameters of interest in the interval | |
| virtual void | SetPdf (RooAbsPdf &pdf) |
| Set the Pdf. | |
| void | SetTestSize (double size) override |
| set the size of the test (rate of Type I error) ( Eg. 0.05 for a 95% Confidence Interval) | |
| double | Size () const override |
| Get the size of the test (eg. rate of Type I error). | |
| void | Streamer (TBuffer &) override |
| void | StreamerNVirtual (TBuffer &ClassDef_StreamerNVirtual_b) |
Static Public Member Functions | |
| static TClass * | Class () |
| static const char * | Class_Name () |
| static constexpr Version_t | Class_Version () |
| static const char * | DeclFileName () |
Protected Member Functions | |
| RooFit::OwningPtr< RooAbsReal > | DoGlobalFit () const |
| perform a global fit | |
| void | DoReset () const |
| clear internal fit result | |
| RooAbsData * | GetData () const |
| RooAbsPdf * | GetPdf () const |
Static Protected Member Functions | |
| static RooFit::OwningPtr< RooFitResult > | DoMinimizeNLL (RooAbsReal *nll) |
| minimize likelihood | |
Protected Attributes | |
| RooArgSet | fAlternateParams |
| RooArgSet specifying alternate parameters for hypothesis test. | |
| RooArgSet | fConditionalObs |
| RooArgSet specifying the conditional observables. | |
| RooAbsData * | fData = nullptr |
| std::unique_ptr< RooFitResult > | fFitResult |
| bool | fGlobalFitDone |
| flag to control if a global fit has been done | |
| RooArgSet | fGlobalObs |
| RooArgSet specifying the global observables. | |
| RooArgSet | fNuisParams |
| RooArgSet specifying nuisance parameters for interval. | |
| RooArgSet | fNullParams |
| RooArgSet specifying null parameters for hypothesis test. | |
| RooAbsPdf * | fPdf = nullptr |
| RooArgSet | fPOI |
| RooArgSet specifying parameters of interest for interval. | |
| double | fSize = 0.0 |
| size of the test (eg. specified rate of Type I error) | |
#include <RooStats/ProfileLikelihoodCalculator.h>
| ProfileLikelihoodCalculator::ProfileLikelihoodCalculator | ( | ) |
Default constructor (needed for I/O).
default constructor
Definition at line 78 of file ProfileLikelihoodCalculator.cxx.
| ProfileLikelihoodCalculator::ProfileLikelihoodCalculator | ( | RooAbsData & | data, |
| RooAbsPdf & | pdf, | ||
| const RooArgSet & | paramsOfInterest, | ||
| double | size = 0.05, | ||
| const RooArgSet * | nullParams = nullptr ) |
Constructor from data, from a full model pdf describing both parameter of interest and nuisance parameters and from the set specifying the parameter of interest (POI).
There is no need to specify the nuisance parameters since they are all other parameters of the model. When using the calculator for performing an hypothesis test one needs to provide also a snapshot (a copy) defining the null parameters and their value. There is no need to pass the alternate parameters. These will be obtained by the value maximizing the likelihood function
Definition at line 80 of file ProfileLikelihoodCalculator.cxx.
| ProfileLikelihoodCalculator::ProfileLikelihoodCalculator | ( | RooAbsData & | data, |
| ModelConfig & | model, | ||
| double | size = 0.05 ) |
Constructor from data and a model configuration If the ModelConfig defines a prior pdf for any of the parameters those will be included as constrained terms in the likelihood function.
Definition at line 89 of file ProfileLikelihoodCalculator.cxx.
|
overridedefault |
destructor cannot delete prod pdf because it will delete all the composing pdf's if (fOwnPdf) delete fPdf; fPdf = 0;
|
static |
|
static |
|
inlinestaticconstexpr |
Definition at line 74 of file ProfileLikelihoodCalculator.h.
|
inlineoverridevirtualinherited |
Get the Confidence level for the test.
Implements RooStats::IntervalCalculator.
Definition at line 103 of file CombinedCalculator.h.
|
inlinestatic |
Definition at line 74 of file ProfileLikelihoodCalculator.h.
|
protected |
perform a global fit
Definition at line 113 of file ProfileLikelihoodCalculator.cxx.
|
staticprotected |
minimize likelihood
Definition at line 156 of file ProfileLikelihoodCalculator.cxx.
|
protected |
clear internal fit result
Definition at line 107 of file ProfileLikelihoodCalculator.cxx.
|
inlineprotectedinherited |
Definition at line 152 of file CombinedCalculator.h.
|
overridevirtual |
Return the hypothesis test result obtained from the likelihood ratio of the maximum likelihood value with the null parameters fixed to their values, with respect to keeping all parameters floating (global maximum likelihood value).
Main interface to get a HypoTestResult.
It does two fits:
The ratio of the likelihood obtained from the conditional MLE to the MLE is the profile likelihood ratio. Wilks' theorem is used to get \(p\)-values.
Implements RooStats::CombinedCalculator.
Definition at line 284 of file ProfileLikelihoodCalculator.cxx.
|
overridevirtual |
Return a likelihood interval.
Main interface to get a RooStats::ConfInterval.
A global fit to the likelihood is performed and the interval is constructed using the profile likelihood ratio function of the POI.
It constructs a profile likelihood ratio, and uses that to construct a RooStats::LikelihoodInterval.
Implements RooStats::CombinedCalculator.
Definition at line 209 of file ProfileLikelihoodCalculator.cxx.
|
inlineprotectedinherited |
Definition at line 151 of file CombinedCalculator.h.
|
inlineoverridevirtual |
Reimplemented from RooStats::CombinedCalculator.
Definition at line 74 of file ProfileLikelihoodCalculator.h.
|
inlineoverridevirtualinherited |
Set the model for the alternate hypothesis.
Implements RooStats::HypoTestCalculator.
Definition at line 122 of file CombinedCalculator.h.
|
inlinevirtualinherited |
set parameter values for the alternate if using a common PDF
Definition at line 140 of file CombinedCalculator.h.
|
inlinevirtualinherited |
Set a common model for both the null and alternate, add to the workspace if not already there.
Definition at line 70 of file HypoTestCalculator.h.
|
inlinevirtualinherited |
set conditional observables needed for computing the NLL
Definition at line 143 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
set the confidence level for the interval (eg. 0.95 for a 95% Confidence Interval)
Implements RooStats::IntervalCalculator.
Definition at line 99 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
Set the DataSet, add to the workspace if not already there.
Implements RooStats::IntervalCalculator.
Definition at line 106 of file CombinedCalculator.h.
|
inlinevirtualinherited |
set global observables needed for computing the NLL
Definition at line 146 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
set the model (in this case can set only the parameters for the null hypothesis)
Implements RooStats::IntervalCalculator.
Definition at line 111 of file CombinedCalculator.h.
|
inlinevirtualinherited |
specify the nuisance parameters (eg. the rest of the parameters)
Definition at line 134 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
Set the model for the null hypothesis.
Implements RooStats::HypoTestCalculator.
Definition at line 120 of file CombinedCalculator.h.
|
inlinevirtualinherited |
set parameter values for the null if using a common PDF
Definition at line 137 of file CombinedCalculator.h.
|
inlinevirtualinherited |
specify the parameters of interest in the interval
Definition at line 131 of file CombinedCalculator.h.
|
inlinevirtualinherited |
Set the Pdf.
Definition at line 128 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
set the size of the test (rate of Type I error) ( Eg. 0.05 for a 95% Confidence Interval)
Implements RooStats::IntervalCalculator.
Definition at line 97 of file CombinedCalculator.h.
|
inlineoverridevirtualinherited |
Get the size of the test (eg. rate of Type I error).
Implements RooStats::IntervalCalculator.
Definition at line 101 of file CombinedCalculator.h.
|
overridevirtual |
Reimplemented from RooStats::CombinedCalculator.
|
inline |
Definition at line 74 of file ProfileLikelihoodCalculator.h.
|
protectedinherited |
RooArgSet specifying alternate parameters for hypothesis test.
Definition at line 160 of file CombinedCalculator.h.
|
protectedinherited |
RooArgSet specifying the conditional observables.
Definition at line 162 of file CombinedCalculator.h.
|
protectedinherited |
Definition at line 157 of file CombinedCalculator.h.
|
mutableprotected |
Definition at line 70 of file ProfileLikelihoodCalculator.h.
|
mutableprotected |
flag to control if a global fit has been done
Definition at line 71 of file ProfileLikelihoodCalculator.h.
|
protectedinherited |
RooArgSet specifying the global observables.
Definition at line 163 of file CombinedCalculator.h.
|
protectedinherited |
RooArgSet specifying nuisance parameters for interval.
Definition at line 161 of file CombinedCalculator.h.
|
protectedinherited |
RooArgSet specifying null parameters for hypothesis test.
Definition at line 159 of file CombinedCalculator.h.
|
protectedinherited |
Definition at line 156 of file CombinedCalculator.h.
|
protectedinherited |
RooArgSet specifying parameters of interest for interval.
Definition at line 158 of file CombinedCalculator.h.
|
protectedinherited |
size of the test (eg. specified rate of Type I error)
Definition at line 154 of file CombinedCalculator.h.